Three-dimensional structures of noncovalent complexes of Citrobacter freundii methionine gamma-lyase with substrates.
Revtovich, S.V., Morozova, E.A., Khurs, E.N., Zakomirdina, L.N., Nikulin, A.D., Demidkina, T.V., Khomutov, R.M.(2011) Biochemistry (Mosc) 76: 564-570
- PubMed: 21639836
- DOI: https://doi.org/10.1134/S0006297911050063
- Primary Citation of Related Structures:
3JW9, 3JWA, 3JWB - PubMed Abstract:
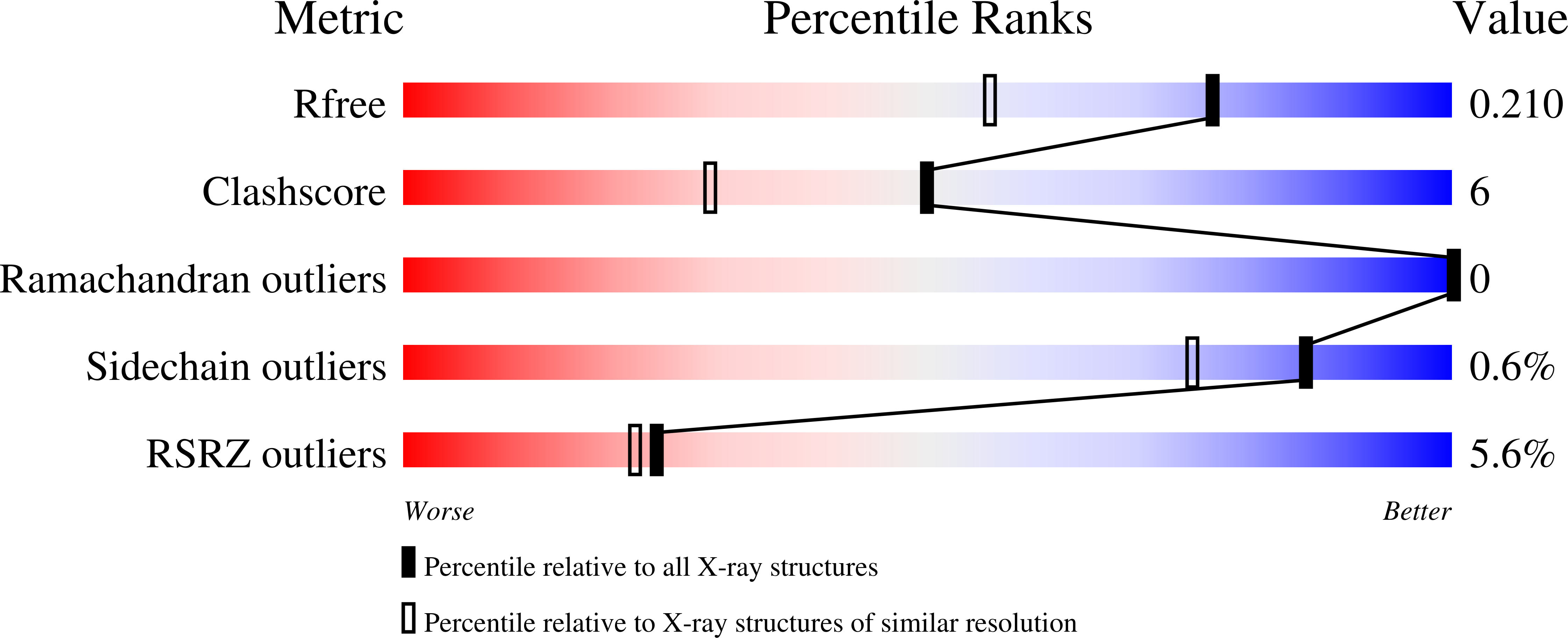
Crystal structures of Citrobacter freundii methionine γ-lyase complexes with the substrates of γ- (L-1-amino-3-methylthiopropylphosphinic acid) and β- (S-ethyl-L-cysteine) elimination reactions and the competitive inhibitor L-norleucine have been determined at 1.45, 1.8, and 1.63 Å resolution, respectively. All three amino acids occupy the active site of the enzyme but do not form a covalent bond with pyridoxal 5'-phosphate. Hydrophobic interactions between the active site residues and the side groups of the substrates and the inhibitor are supposed to cause noncovalent binding. Arg374 and Ser339 are involved in the binding of carboxyl groups of the substrates and the inhibitor. The hydroxyl of Tyr113 is a potential acceptor of a proton from the amino groups of the amino acids.
Organizational Affiliation:
Engelhardt Institute of Molecular Biology, Russian Academy of Sciences, Moscow. [email protected]