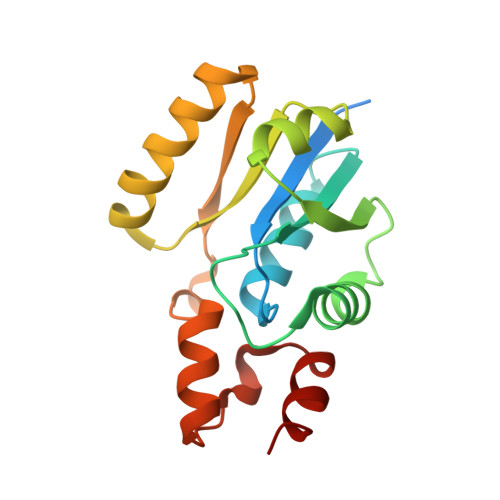
Kinetic, thermodynamic, and structural insight into the mechanism of phosphopantetheine adenylyltransferase from Mycobacterium tuberculosis.
Wubben, T.J., Mesecar, A.D.(2010) J Mol Biol 404: 202-219
- PubMed: 20851704
- DOI: https://doi.org/10.1016/j.jmb.2010.09.002
- Primary Citation of Related Structures:
3NBA, 3NBK - PubMed Abstract:
Phosphopantetheine adenylyltransferase (PPAT) catalyzes the penultimate step in the coenzyme A (CoA) biosynthetic pathway, reversibly transferring an adenylyl group from ATP to 4'-phosphopantetheine (PhP) to form dephosphocoenzyme A. This reaction sits at the branch point between the de novo pathway and the salvage pathway, and has been shown to be a rate-limiting step in the biosynthesis of CoA. Importantly, bacterial and mammalian PPATs share little sequence homology, making the enzyme a potential target for antibiotic development. A series of steady-state kinetic, product inhibition, and direct binding studies with Mycobacterium tuberculosis PPAT (MtPPAT) was conducted and suggests that the enzyme utilizes a nonrapid-equilibrium random bi-bi mechanism. The kinetic response of MtPPAT to the binding of ATP was observed to be sigmoidal under fixed PhP concentrations, but substrate inhibition was observed at high PhP concentrations under subsaturating ATP concentrations, suggesting a preferred pathway to ternary complex formation. Negative cooperativity in the kinetic response of MtPPAT to PhP binding was observed under certain conditions and confirmed thermodynamically by isothermal titration calorimetry, suggesting the formation of an asymmetric quaternary structure during sequential ligation of substrates. Asymmetry in binding was also observed in isothermal titration calorimetry experiments with dephosphocoenzyme A and CoA. X-ray structures of MtPPAT in complex with PhP and the nonhydrolyzable ATP analogue adenosine-5'-[(α,β)-methyleno]triphosphate were solved to 1.57 Å and 2.68 Å, respectively. These crystal structures reveal small conformational changes in enzyme structure upon ligand binding, which may play a role in the nonrapid-equilibrium mechanism. We suggest that the proposed kinetic mechanism and asymmetric character in MtPPAT ligand binding may provide a means of reaction and pathway regulation in addition to that of the previously determined CoA feedback.
Organizational Affiliation:
Department of Biochemistry and Molecular Genetics, University of Illinois at Chicago, Chicago, IL 60607, USA.