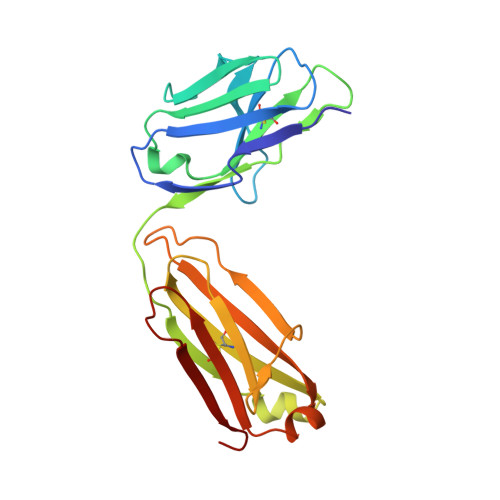
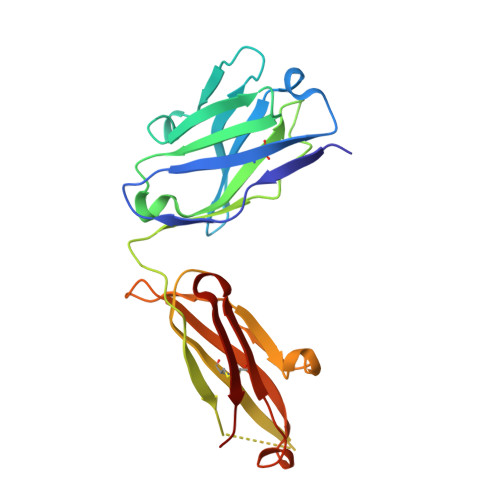
Structural basis of immunosuppression by the therapeutic antibody daclizumab
Yang, H., Wang, J., Du, J., Zhong, C., Zhang, D., Guo, H., Guo, Y., Ding, J.(2010) Cell Res 20: 1361-1371
- PubMed: 20820193
- DOI: https://doi.org/10.1038/cr.2010.130
- Primary Citation of Related Structures:
3NFP, 3NFS - PubMed Abstract:
Interleukin-2 (IL)-2 signaling plays a pivotal role in the activation of immune responses, and drugs that block this pathway have been shown to be effective for the immunosuppression in patients with organ transplantation to alleviate/eliminate allograft rejection. The first humanized monoclonal antibody (mAb) daclizumab falls into this category and shows high specificity and affinity against a key component of the IL-2 receptor complex, namely IL-2Rα. To reveal the molecular mechanism of the inhibition of the IL-2 signaling pathway by daclizumab, we determined the crystal structures of the daclizumab Fab in free form and in complex with the IL-2Rα ectodomain at 2.6 and 2.8 Å resolution, respectively. The daclizumab Fab adopts a similar conformation in the presence or absence of the IL-2Rα ectodomain. The antigen-binding site of daclizumab is mainly composed of five complementarity determining regions (CDRs) that form a large positively charged surface depression and two flanking patches that are generally hydrophobic. The conformational epitope consists of several discontinuous segments of the IL-2Rα ectodomain, a large portion of which overlaps with the regions that interact with IL-2, suggesting that the binding of daclizumab to IL-2Rα would prevent the IL-2 binding to IL-2Rα and the subsequent formation of the IL-2/IL-2Rαβγ(c) complex, and therefore block the IL-2 signaling pathway. These results also have implications for the design and development of improved mAb drugs targeting IL-2Rα.
Organizational Affiliation:
State Key Laboratory of Molecular Biology and Research Center for Structural Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China.