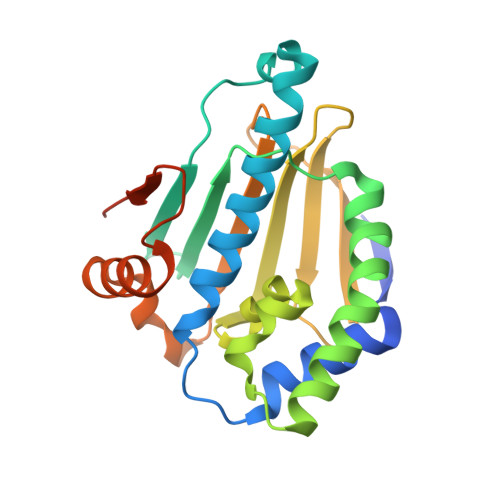
Design and SAR of macrocyclic Hsp90 inhibitors with increased metabolic stability and potent cell-proliferation activity.
Zapf, C.W., Bloom, J.D., McBean, J.L., Dushin, R.G., Nittoli, T., Ingalls, C., Sutherland, A.G., Sonye, J.P., Eid, C.N., Golas, J., Liu, H., Boschelli, F., Hu, Y., Vogan, E., Levin, J.I.(2011) Bioorg Med Chem Lett 21: 2278-2282
- PubMed: 21420297
- DOI: https://doi.org/10.1016/j.bmcl.2011.02.101
- Primary Citation of Related Structures:
3QTF - PubMed Abstract:
A novel series of macrocyclic ortho-aminobenzamide Hsp90 inhibitors is reported. A basic nitrogen within the tether linking the aniline nitrogen atom to a tetrahydroindolone moiety allowed access to compounds with good physical properties. Important structure-activity relationship information was obtained from this series which led to the discovery of a soluble and stable compound which is potent in an Hsp90 binding and cell-proliferation assay.
Organizational Affiliation:
Medicinal Chemistry, Pfizer, 401 N. Middletown Road, Pearl River, NY 10965, USA. [email protected]