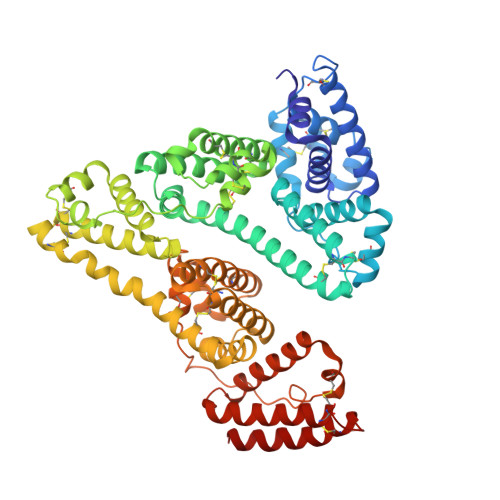
Large-scale production of functional human serum albumin from transgenic rice seeds.
He, Y., Ning, T., Xie, T., Qiu, Q., Zhang, L., Sun, Y., Jiang, D., Fu, K., Yin, F., Zhang, W., Shen, L., Wang, H., Li, J., Lin, Q., Sun, Y., Li, H., Zhu, Y., Yang, D.(2011) Proc Natl Acad Sci U S A 108: 19078-19083
- PubMed: 22042856
- DOI: https://doi.org/10.1073/pnas.1109736108
- Primary Citation of Related Structures:
3SQJ - PubMed Abstract:
Human serum albumin (HSA) is widely used in clinical and cell culture applications. Conventional production of HSA from human blood is limited by the availability of blood donation and the high risk of viral transmission from donors. Here, we report the production of Oryza sativa recombinant HSA (OsrHSA) from transgenic rice seeds. The level of OsrHSA reached 10.58% of the total soluble protein of the rice grain. Large-scale production of OsrHSA generated protein with a purity >99% and a productivity rate of 2.75 g/kg brown rice. Physical and biochemical characterization of OsrHSA revealed it to be equivalent to plasma-derived HSA (pHSA). The efficiency of OsrHSA in promoting cell growth and treating liver cirrhosis in rats was similar to that of pHSA. Furthermore, OsrHSA displays similar in vitro and in vivo immunogenicity as pHSA. Our results suggest that a rice seed bioreactor produces cost-effective recombinant HSA that is safe and can help to satisfy an increasing worldwide demand for human serum albumin.
Organizational Affiliation:
Engineering Research Center for Plant Biotechnology and Germplasm Utilization, Ministry of Education, State Key Laboratory of Hybrid Rice, College of Life Sciences, Wuhan University, Wuhan 430072, China.