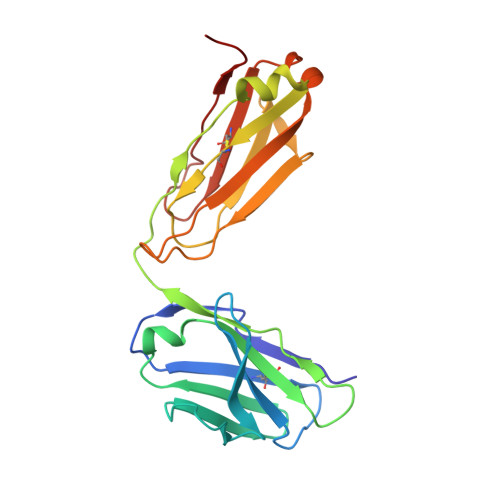
Structure-Guided Design of a High-Affinity Platelet Integrin alphaIIbbeta3 Receptor Antagonist That Disrupts Mg2+ Binding to the MIDAS
Zhu, J., Choi, W.S., McCoy, J.G., Negri, A., Zhu, J., Naini, S., Li, J., Shen, M., Huang, W., Bougie, D., Rasmussen, M., Aster, R., Thomas, C.J., Filizola, M., Springer, T.A., Coller, B.S.(2012) Sci Transl Med 4: 125ra32-125ra32
- PubMed: 22422993
- DOI: https://doi.org/10.1126/scitranslmed.3003576
- Primary Citation of Related Structures:
3T3M, 3T3P - PubMed Abstract:
An integrin found on platelets, α(IIb)β(3) mediates platelet aggregation, and α(IIb)β(3) antagonists are effective antithrombotic agents in the clinic. Ligands bind to integrins in part by coordinating a magnesium ion (Mg(2+)) located in the β subunit metal ion-dependent adhesion site (MIDAS). Drugs patterned on the integrin ligand sequence Arg-Gly-Asp have a basic moiety that binds the α(IIb) subunit and a carboxyl group that coordinates the MIDAS Mg(2+) in the β(3) subunits. They induce conformational changes in the β(3) subunit that may have negative consequences such as exposing previously hidden epitopes and inducing the active conformation of the receptor. We recently reported an inhibitor of α(IIb)β(3) (RUC-1) that binds exclusively to the α(IIb) subunit; here, we report the structure-based design and synthesis of RUC-2, a RUC-1 derivative with a ~100-fold higher affinity. RUC-2 does not induce major conformational changes in β(3) as judged by monoclonal antibody binding, light scattering, gel chromatography, electron microscopy, and a receptor priming assay. X-ray crystallography of the RUC-2-α(IIb)β(3) headpiece complex in 1 mM calcium ion (Ca(2+))/5 mM Mg(2+) at 2.6 Å revealed that RUC-2 binds to α(IIb) the way RUC-1 does, but in addition, it binds to the β(3) MIDAS residue glutamic acid 220, thus displacing Mg(2+) from the MIDAS. When the Mg(2+) concentration was increased to 20 mM, however, Mg(2+) was identified in the MIDAS and RUC-2 was absent. RUC-2's ability to inhibit ligand binding and platelet aggregation was diminished by increasing the Mg(2+) concentration. Thus, RUC-2 inhibits ligand binding by a mechanism different from that of all other α(IIb)β(3) antagonists and may offer advantages as a therapeutic agent.
Organizational Affiliation:
Immune Disease Institute, Children's Hospital Boston, and Department of Pathology, Harvard Medical School, Boston, MA 02115, USA.