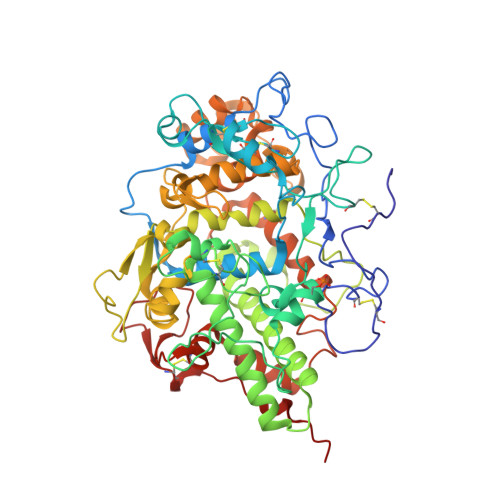
Bovine carbonyl lactoperoxidase structure at 2.0 angstrom resolution and infrared spectra as a function of pH.
Singh, A.K., Smith, M.L., Yamini, S., Ohlsson, P.I., Sinha, M., Kaur, P., Sharma, S., Paul, J.A., Singh, T.P., Paul, K.G.(2012) Protein J 31: 598-608
- PubMed: 22886082
- DOI: https://doi.org/10.1007/s10930-012-9436-3
- Primary Citation of Related Structures:
3V6Q - PubMed Abstract:
Lactoperoxidase (LPO) is a hemeprotein catalyzing the oxidation of thiocyanate and I(-) into antimicrobials and small aromatic organics after being itself oxidized by H(2)O(2). LPO is excreted by the lungs, mammary glands, found in saliva and tears and protects mammals against bacterial, fungal and viral invasion. The Fe(II) form binds CO which inactivates LPO like many other hemeproteins. We present the 3-dimensional structure of CO-LPO at 2.0Å resolution and infrared (IR) spectra of the iron-bound CO stretch from pH 3 to 8.8 at 1 cm(-1) resolution. The observed Fe-C-O bond angle of 132° is more acute than the electronically related Fe(III), CN-LPO with a Fe-C-N angle of 161°. The orientations of the two ligands are different with the oxygen of CO pointing towards the imidazole of distal His109 while the nitrogen of CN points away, the Fe(II) moves towards His109 while the Fe(III) moves away; both movements are consistent with a hydrogen bond between the distal His109 and CO, but not to the nitrogen of CN-LPO. The IR spectra of CO-LPO exhibit two major CO absorbances with pH dependent relative intensities. Both crystallographic and IR data suggest proton donation to the CO oxygen by His109 with a pK ≈ 4; close to the pH of greatest enzyme turnover. The IR absorbance maxima are consistent with a first order correlation between frequency and Fe(III)/Fe(II) reduction potential at pH 7; both band widths at half-height correlate with electron density donation from Fe(II) to CO as gauged by the reduction potential.
Organizational Affiliation:
Department of Biophysics, All India Institute of Medical Sciences, New Delhi, 110 029, India. [email protected]