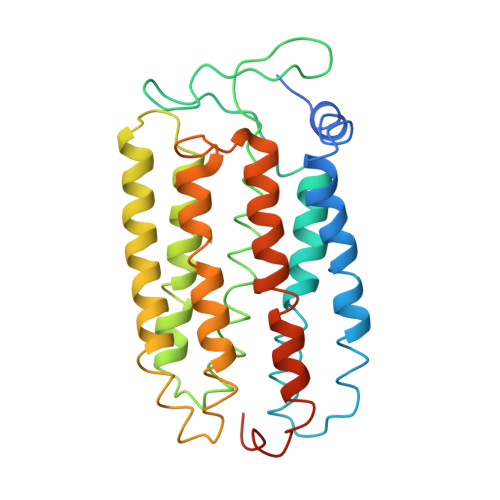
Large deformation of helix F during the photoreaction cycle of Pharaonis halorhodopsin in complex with azide
Nakanishi, T., Kanada, S., Murakami, M., Ihara, K., Kouyama, T.(2013) Biophys J 104: 377-385
- PubMed: 23442859
- DOI: https://doi.org/10.1016/j.bpj.2012.12.018
- Primary Citation of Related Structures:
3VVK - PubMed Abstract:
Halorhodopsin from Natronomonas pharaonis (pHR), a retinylidene protein that functions as a light-driven chloride ion pump, is converted into a proton pump in the presence of azide ion. To clarify this conversion, we investigated light-induced structural changes in pHR using a C2 crystal that was prepared in the presence of Cl(-) and subsequently soaked in a solution containing azide ion. When the pHR-azide complex was illuminated at pH 9, a profound outward movement (∼4 Å) of the cytoplasmic half of helix F was observed in a subunit with the EF loop facing an open space. This movement created a long water channel between the retinal Schiff base and the cytoplasmic surface, along which a proton could be transported. Meanwhile, the middle moiety of helix C moved inward, leading to shrinkage of the primary anion-binding site (site I), and the azide molecule in site I was expelled out to the extracellular medium. The results suggest that the cytoplasmic half of helix F and the middle moiety of helix C act as different types of valves for active proton transport.
Organizational Affiliation:
Graduate School of Science, Nagoya University, Nagoya, Japan.