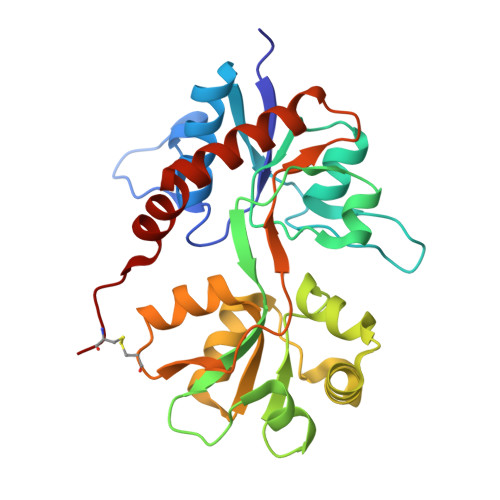
Structural and pharmacological characterization of phenylalanine-based AMPA receptor antagonists at kainate receptors
Venskutonyte, R., Frydenvang, K., Valades, E.A., Szymanska, E., Johansen, T.N., Kastrup, J.S., Pickering, D.S.(2012) ChemMedChem 7: 1793-1798
- PubMed: 22407805
- DOI: https://doi.org/10.1002/cmdc.201100599
- Primary Citation of Related Structures:
4DLD - PubMed Abstract:
Continued efforts into the discovery of ligands that target ionotropic glutamate receptors (iGluRs) are important for studies of the physiological roles of the various iGluR subtypes as well as for the search for drugs that can be used in the treatment of diseases of the central nervous system. A new series of phenylalanine derivatives that target iGluRs was reported to bind AMPA receptors. Herein we report our studies of these compounds at the kainate receptors GluK1-3. Several compounds bind with micromolar affinity at GluK1 and GluK3, but do not bind GluK2. The crystal structure of the most potent compound in the ligand binding domain of GluK1 revealed different modes of binding to GluK1 and GluA2, due primarily to residues Ser741 (GluK1) and Met729 (GluA2). The compound was shown to be slightly more potent at GluK1 than at AMPA receptors and to induce a domain closure similar to that observed in GluK1 structures with partial agonists.
Organizational Affiliation:
Department Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen, Universitetsparken 2, 2100 Copenhagen, Denmark.