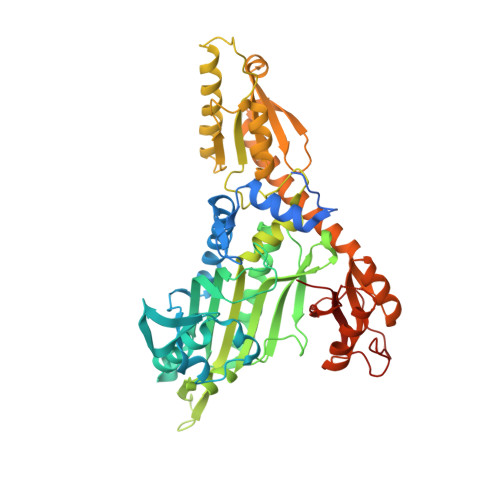
Conformational changes in human prolyl-tRNA synthetase upon binding of the substrates proline and ATP and the inhibitor halofuginone.
Son, J., Lee, E.H., Park, M., Kim, J.H., Kim, J., Kim, S., Jeon, Y.H., Hwang, K.Y.(2013) Acta Crystallogr D Biol Crystallogr 69: 2136-2145
- PubMed: 24100331
- DOI: https://doi.org/10.1107/S0907444913020556
- Primary Citation of Related Structures:
4K86, 4K87, 4K88 - PubMed Abstract:
Aminoacyl-tRNA synthetases recognize cognate amino acids and tRNAs from their noncognate counterparts and catalyze the formation of aminoacyl-tRNAs. Halofuginone (HF), a coccidiostat used in veterinary medicine, exerts its effects by acting as a high-affinity inhibitor of the enzyme glutamyl-prolyl-tRNA synthetase (EPRS). In order to elucidate the precise molecular basis of this inhibition mechanism of human EPRS, the crystal structures of the prolyl-tRNA synthetase domain of human EPRS (hPRS) at 2.4 Å resolution (hPRS-apo), of hPRS complexed with ATP and the substrate proline at 2.3 Å resolution (hPRS-sub) and of hPRS complexed with HF at 2.62 Å resolution (hPRS-HF) are presented. These structures show plainly that motif 1 functions as a cap in hPRS, which is loosely opened in hPRS-apo, tightly closed in hPRS-sub and incorrectly closed in hPRS-HF. In addition, the structural analyses are consistent with more effective binding of hPRS to HF with ATP. Mutagenesis and biochemical analysis confirmed the key roles of two residues, Phe1097 and Arg1152, in the HF inhibition mechanism. These structures will lead to the development of more potent and selective hPRS inhibitors for promoting inflammatory resolution.
Organizational Affiliation:
Department of Biosystems and Biotechnology, Korea University, Anam-dong 5, Seongbuk-gu, Seoul 136-701, Republic of Korea.