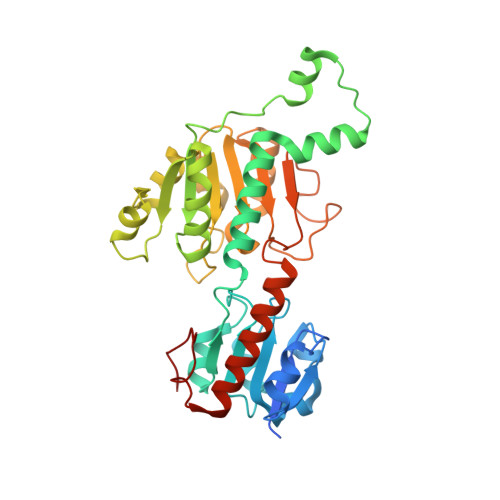
Crystal structures of human CtBP in complex with substrate MTOB reveal active site features useful for inhibitor design.
Hilbert, B.J., Grossmann, S.R., Schiffer, C.A., Royer, W.E.(2014) FEBS Lett 588: 1743-1748
- PubMed: 24657618
- DOI: https://doi.org/10.1016/j.febslet.2014.03.026
- Primary Citation of Related Structures:
4LCE, 4LCJ - PubMed Abstract:
The oncogenic corepressors C-terminal Binding Protein (CtBP) 1 and 2 harbor regulatory d-isomer specific 2-hydroxyacid dehydrogenase (d2-HDH) domains. 4-Methylthio 2-oxobutyric acid (MTOB) exhibits substrate inhibition and can interfere with CtBP oncogenic activity in cell culture and mice. Crystal structures of human CtBP1 and CtBP2 in complex with MTOB and NAD(+) revealed two key features: a conserved tryptophan that likely contributes to substrate specificity and a hydrophilic cavity that links MTOB with an NAD(+) phosphate. Neither feature is present in other d2-HDH enzymes. These structures thus offer key opportunities for the development of highly selective anti-neoplastic CtBP inhibitors.
Organizational Affiliation:
Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, MA 01605, USA.