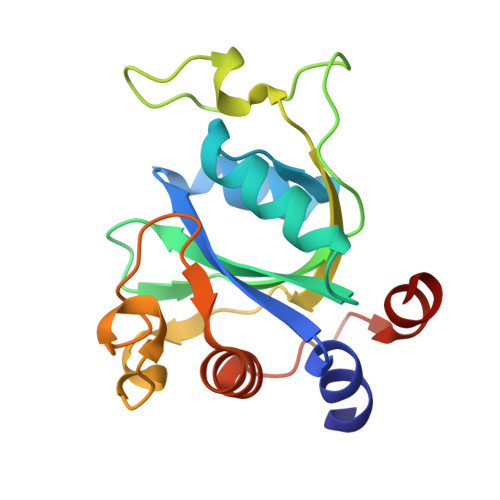
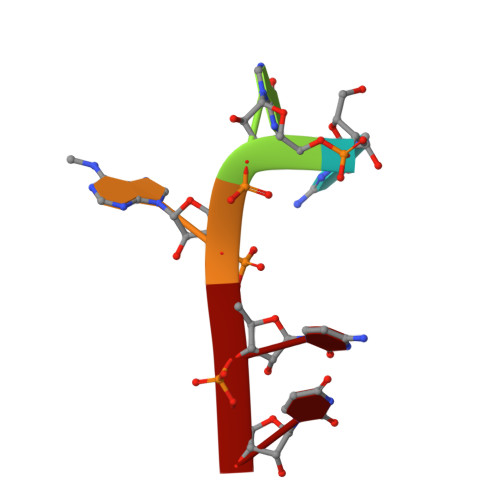
Structural basis for selective binding of m(6)A RNA by the YTHDC1 YTH domain.
Xu, C., Wang, X., Liu, K., Roundtree, I.A., Tempel, W., Li, Y., Lu, Z., He, C., Min, J.(2014) Nat Chem Biol 10: 927-929
- PubMed: 25242552
- DOI: https://doi.org/10.1038/nchembio.1654
- Primary Citation of Related Structures:
4R3H, 4R3I - PubMed Abstract:
N(6)-methyladenosine (m(6)A) is the most abundant internal modification of nearly all eukaryotic mRNAs and has recently been reported to be recognized by the YTH domain family proteins. Here we present the crystal structures of the YTH domain of YTHDC1, a member of the YTH domain family, and its complex with an m(6)A-containing RNA. Our structural studies, together with transcriptome-wide identification of YTHDC1-binding sites and biochemical experiments, not only reveal the specific mode of m(6)A-YTH binding but also explain the preferential recognition of the GG(m(6)A)C sequences by YTHDC1.
Organizational Affiliation:
1] Structural Genomics Consortium, University of Toronto, Toronto, Ontario, Canada. [2]. [email protected].