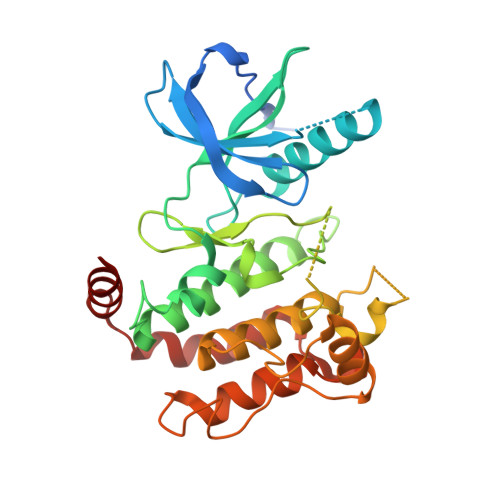
Crystal Structures of Human Rip2 Kinase Catalytic Domain Complexed with ATP-Competitive Inhibitors: Foundations for Understanding Inhibitor Selectivity.
Charnley, A.K., Convery, M.A., Lakdawala Shah, A., Jones, E., Hardwicke, P., Bridges, A., Ouellette, M., Totoritis, R., Schwartz, B., King, B.W., Wisnoski, D.D., Kang, J., Eidam, P.M., Votta, B.J., Gough, P.J., Marquis, R.W., Bertin, J., Casillas, L.(2015) Bioorg Med Chem 23: 7000
- PubMed: 26455654
- DOI: https://doi.org/10.1016/j.bmc.2015.09.038
- Primary Citation of Related Structures:
5AR2, 5AR3, 5AR4, 5AR5, 5AR7, 5AR8 - PubMed Abstract:
Receptor interacting protein 2 (RIP2) is an intracellular kinase and key signaling partner for the pattern recognition receptors NOD1 and NOD2 (nucleotide-binding oligomerization domain-containing proteins 1 and 2). As such, RIP2 represents an attractive target to probe the role of these pathways in disease. In an effort to design potent and selective inhibitors of RIP2 we established a crystallographic system and determined the structure of the RIP2 kinase domain in an apo form and also in complex with multiple inhibitors including AMP-PCP (β,γ-Methyleneadenosine 5'-triphosphate, a non-hydrolysable adenosine triphosphate mimic) and structurally diverse ATP competitive chemotypes identified via a high-throughput screening campaign. These structures represent the first set of diverse RIP2-inhibitor co-crystal structures and demonstrate that the protein possesses the ability to adopt multiple DFG-in as well as DFG-out and C-helix out conformations. These structures reveal key protein-inhibitor structural insights and serve as the foundation for establishing a robust structure-based drug design effort to identify both potent and highly selective inhibitors of RIP2 kinase.
Organizational Affiliation:
Pattern Recognition Receptor Discovery Performance Unit, Immuno-Inflammation Therapy Area, GlaxoSmithKline, 1250 S. Collegeville Road, Collegeville, PA 19426, USA. Electronic address: [email protected].