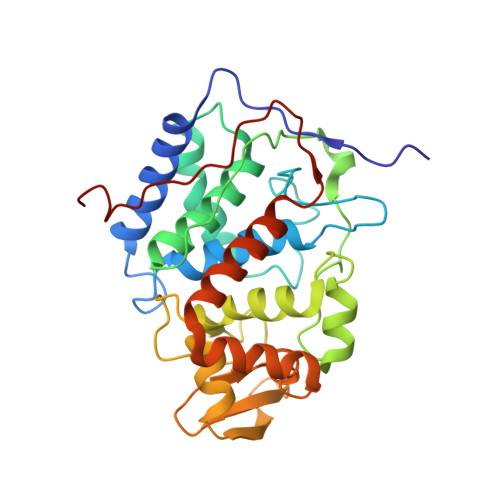
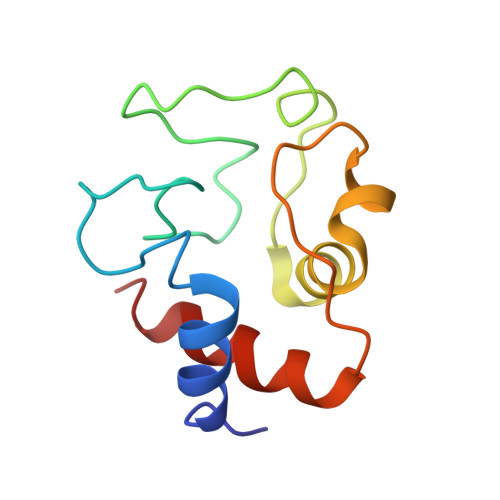
Constraints on the Radical Cation Center of Cytochrome c Peroxidase for Electron Transfer from Cytochrome c.
Payne, T.M., Yee, E.F., Dzikovski, B., Crane, B.R.(2016) Biochemistry 55: 4807-4822
- PubMed: 27499202
- DOI: https://doi.org/10.1021/acs.biochem.6b00262
- Primary Citation of Related Structures:
5CIB, 5CIC, 5CID, 5CIE, 5CIF, 5CIG, 5CIH - PubMed Abstract:
The tryptophan 191 cation radical of cytochrome c peroxidase (CcP) compound I (Cpd I) mediates long-range electron transfer (ET) to cytochrome c (Cc). Here we test the effects of chemical substitution at position 191. CcP W191Y forms a stable tyrosyl radical upon reaction with peroxide and produces spectral properties similar to those of Cpd I but has low reactivity toward reduced Cc. CcP W191G and W191F variants also have low activity, as do redox ligands that bind within the W191G cavity. Crystal structures of complexes between Cc and CcP W191X (X = Y, F, or G), as well as W191G with four bound ligands reveal similar 1:1 association modes and heme pocket conformations. The ligands display structural disorder in the pocket and do not hydrogen bond to Asp235, as does Trp191. Well-ordered Tyr191 directs its hydroxyl group toward the porphyrin ring, with no basic residue in the range of interaction. CcP W191X (X = Y, F, or G) variants substituted with zinc-porphyrin (ZnP) undergo photoinduced ET with Cc(III). Their slow charge recombination kinetics that result from loss of the radical center allow resolution of difference spectra for the charge-separated state [ZnP(+), Cc(II)]. The change from a phenyl moiety at position 191 in W191F to a water-filled cavity in W191G produces effects on ET rates much weaker than the effects of the change from Trp to Phe. Low net reactivity of W191Y toward Cc(II) derives either from the inability of ZnP(+) or the Fe-CcP ferryl to oxidize Tyr or from the low potential of the resulting neutral Tyr radical.
Organizational Affiliation:
Department of Chemistry and Chemical Biology, Cornell University , Ithaca, New York 14853, United States.