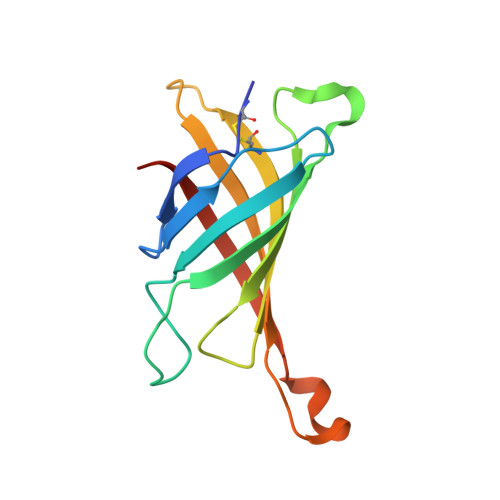
Structural Characterization of the Avidin Interactions with Fluorescent Pyrene-Conjugates: 1-Biotinylpyrene and 1-Desthiobiotinylpyrene.
Strzelczyk, P., Plazuk, D., Zakrzewski, J., Bujacz, G.(2016) Molecules 21
- PubMed: 27689976
- DOI: https://doi.org/10.3390/molecules21101270
- Primary Citation of Related Structures:
5IRU, 5IRW - PubMed Abstract:
Avidin is a tetrameric protein that belongs to the calycin superfamily. It has been studied mainly because of its extraordinary affinity to biotin, which led to a wide range of applications based on the avidin-biotin system. In the present study, we report the first crystal structures of avidin in a complex with two novel fluorescent pyrene derivatives: 1-biotinylpyrene (B9P) and 1-desthiobiotinylpyrene (D9P). The crystal structures were solved by molecular replacement using the coordinates of avidin molecule as a starting model and the final models of avidin/B9P and avidin/D9P were refined to resolutions of 2.0 Å and 2.1 Å, respectively. Our data reveal changes in loop conformation as well as in overall fold and quaternary arrangement of the avidin upon the binding of these fluorescent probes. Moreover, the crystal structures allowed analysis of the details of the interactions between the protein and the pyrene derivatives. Structural description of the complexes will contribute to the design of conjugates for expanding the capabilities of avidin-biotin technology.
Organizational Affiliation:
Institute of Technical Biochemistry, Faculty of Biotechnology and Food Sciences, Lodz University of Technology, 90-924 Łódź, Stefanowskiego 4/10, Poland. [email protected].