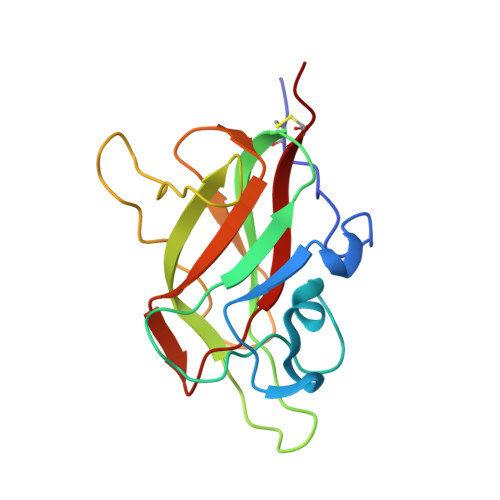
Architecture and hydration of the arginine-binding site of neuropilin-1.
Mota, F., Fotinou, C., Rana, R.R., Chan, A.W.E., Yelland, T., Arooz, M.T., O'Leary, A.P., Hutton, J., Frankel, P., Zachary, I., Selwood, D., Djordjevic, S.(2018) FEBS J 285: 1290-1304
- PubMed: 29430837
- DOI: https://doi.org/10.1111/febs.14405
- Primary Citation of Related Structures:
5IJR, 5IYY, 5J1X, 5JGI, 5JGQ, 5JHK - PubMed Abstract:
Neuropilin-1 (NRP1) is a transmembrane co-receptor involved in binding interactions with variety of ligands and receptors, including receptor tyrosine kinases. Expression of NRP1 in several cancers correlates with cancer stages and poor prognosis. Thus, NRP1 has been considered a therapeutic target and is the focus of multiple drug discovery initiatives. Vascular endothelial growth factor (VEGF) binds to the b1 domain of NRP1 through interactions between the C-terminal arginine of VEGF and residues in the NRP1-binding site including Tyr297, Tyr353, Asp320, Ser346 and Thr349. We obtained several complexes of the synthetic ligands and the NRP1-b1 domain and used X-ray crystallography and computational methods to analyse atomic details and hydration profile of this binding site. We observed side chain flexibility for Tyr297 and Asp320 in the six new high-resolution crystal structures of arginine analogues bound to NRP1. In addition, we identified conserved water molecules in binding site regions which can be targeted for drug design. The computational prediction of the VEGF ligand-binding site hydration map of NRP1 was in agreement with the experimentally derived, conserved hydration structure. Displacement of certain conserved water molecules by a ligand's functional groups may contribute to binding affinity, whilst other water molecules perform as protein-ligand bridges. Our report provides a comprehensive description of the binding site for the peptidic ligands' C-terminal arginines in the b1 domain of NRP1, highlights the importance of conserved structural waters in drug design and validates the utility of the computational hydration map prediction method in the context of neuropilin. The structures were deposited to the PDB with accession numbers PDB ID: 5IJR, 5IYY, 5JHK, 5J1X, 5JGQ, 5JGI.
Organizational Affiliation:
Magnus Life, Magnus Life Science, London, UK.