Chroman-4-One Derivatives Targeting Pteridine Reductase 1 and Showing Anti-Parasitic Activity.
Di Pisa, F., Landi, G., Dello Iacono, L., Pozzi, C., Borsari, C., Ferrari, S., Santucci, M., Santarem, N., Cordeiro-da-Silva, A., Moraes, C.B., Alcantara, L.M., Fontana, V., Freitas-Junior, L.H., Gul, S., Kuzikov, M., Behrens, B., Pohner, I., Wade, R.C., Costi, M.P., Mangani, S.(2017) Molecules 22
- PubMed: 28282886
- DOI: https://doi.org/10.3390/molecules22030426
- Primary Citation of Related Structures:
5K6A, 5L42, 5L4N - PubMed Abstract:
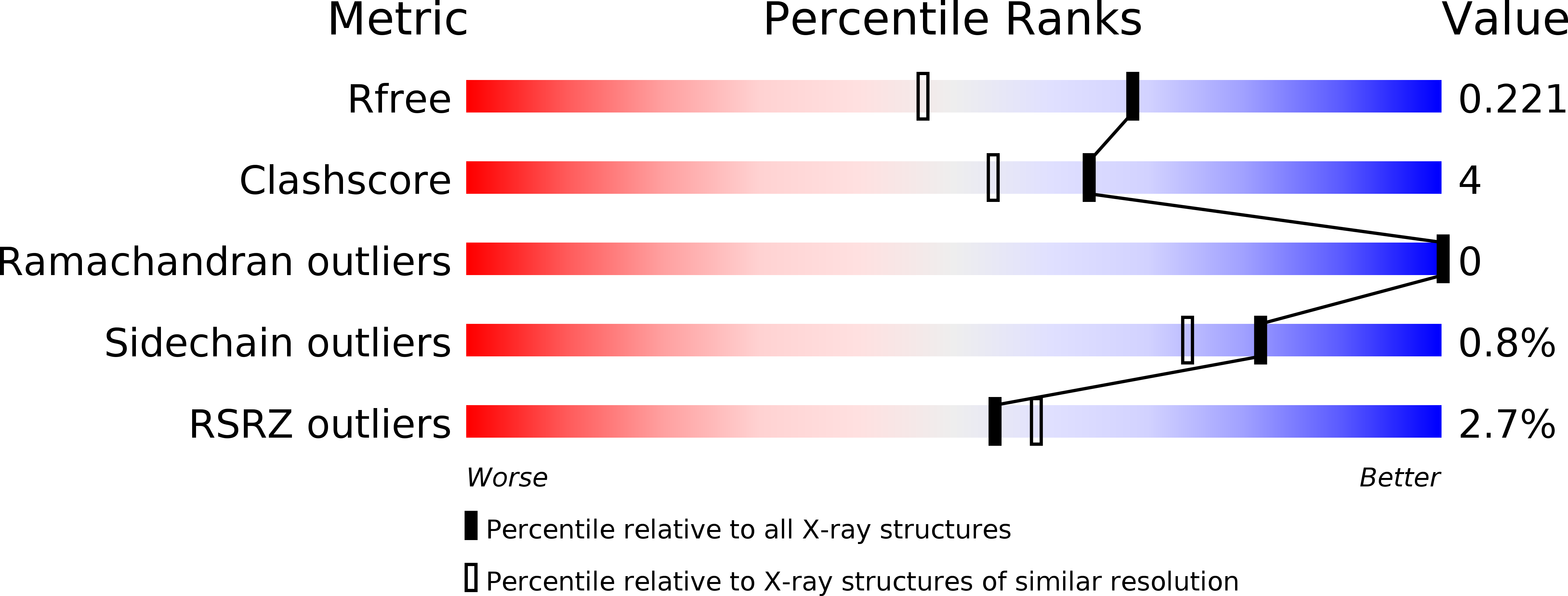
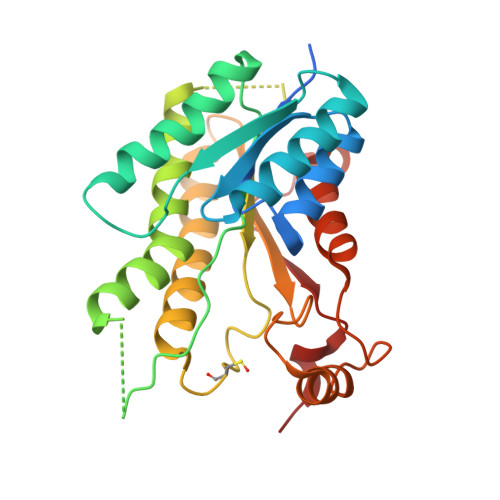
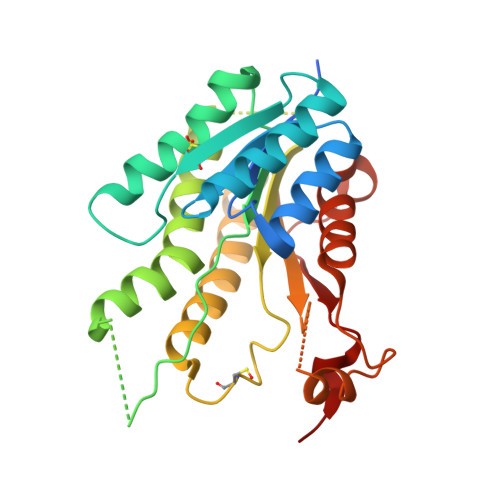
Flavonoids have previously been identified as antiparasitic agents and pteridine reductase 1 (PTR1) inhibitors. Herein, we focus our attention on the chroman-4-one scaffold. Three chroman-4-one analogues ( 1 - 3 ) of previously published chromen-4-one derivatives were synthesized and biologically evaluated against parasitic enzymes ( Trypanosoma brucei PTR1- Tb PTR1 and Leishmania major-Lm PTR1) and parasites ( Trypanosoma brucei and Leishmania infantum ). A crystal structure of Tb PTR1 in complex with compound 1 and the first crystal structures of Lm PTR1-flavanone complexes (compounds 1 and 3 ) were solved. The inhibitory activity of the chroman-4-one and chromen-4-one derivatives was explained by comparison of observed and predicted binding modes of the compounds. Compound 1 showed activity both against the targeted enzymes and the parasites with a selectivity index greater than 7 and a low toxicity. Our results provide a basis for further scaffold optimization and structure-based drug design aimed at the identification of potent anti-trypanosomatidic compounds targeting multiple PTR1 variants.
Organizational Affiliation:
Department of Biotechnology, Chemistry and Pharmacy, University of Siena, 53100 Siena, Italy. [email protected].