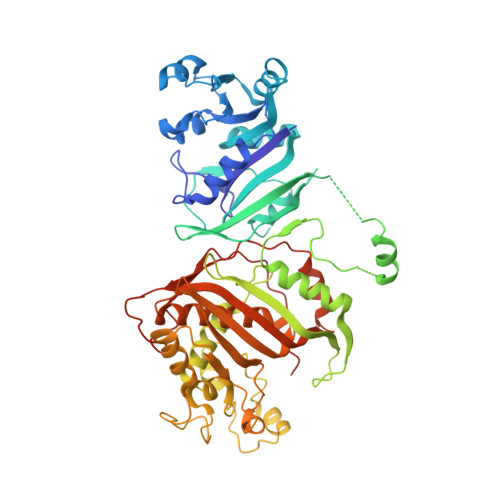
Discovery of Potent and Selective Leads against Toxoplasma gondii Dihydrofolate Reductase via Structure-Based Design.
Welsch, M.E., Zhou, J., Gao, Y., Yan, Y., Porter, G., Agnihotri, G., Li, Y., Lu, H., Chen, Z., Thomas, S.B.(2016) ACS Med Chem Lett 7: 1124-1129
- PubMed: 27994750
- DOI: https://doi.org/10.1021/acsmedchemlett.6b00328
- Primary Citation of Related Structures:
5T0L - PubMed Abstract:
Current treatment of toxoplasmosis targets the parasite's folate metabolism through inhibition of dihydrofolate reductase (DHFR). The most widely used DHFR antagonist, pyrimethamine, was introduced over 60 years ago and is associated with toxicity that can be largely attributed to a similar affinity for parasite and human DHFR. Computational analysis of biochemical differences between Toxoplasma gondii and human DHFR enabled the design of inhibitors with both improved potency and selectivity. The approach described herein yielded TRC-19, a promising lead with an IC 50 of 9 nM and 89-fold selectivity in favor of Toxoplasma gondii DHFR, as well as crystallographic data to substantiate in silico methodology. Overall, 50% of synthesized in silico designs met hit threshold criteria of IC 50 < 10 μM and >2-fold selectivity favoring Toxoplasma gondii , further demonstrating the efficiency of our structure-based drug design approach.
Organizational Affiliation:
Turing Pharmaceuticals AG , Research & Development, 1177 Avenue of the Americas, 39th Floor, New York, New York 10036, United States.