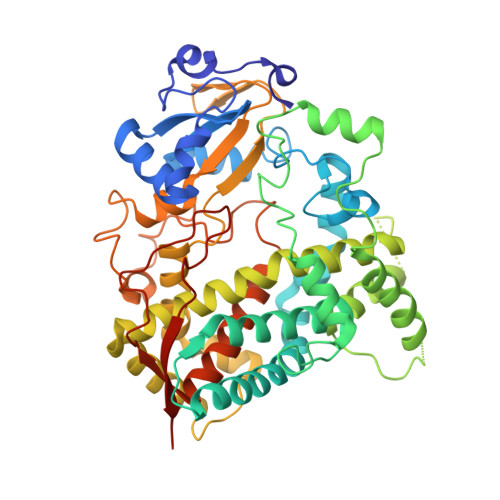
Structural basis for regiospecific midazolam oxidation by human cytochrome P450 3A4.
Sevrioukova, I.F., Poulos, T.L.(2017) Proc Natl Acad Sci U S A 114: 486-491
- PubMed: 28031486
- DOI: https://doi.org/10.1073/pnas.1616198114
- Primary Citation of Related Structures:
5TE8 - PubMed Abstract:
Human cytochrome P450 3A4 (CYP3A4) is a major hepatic and intestinal enzyme that oxidizes more than 60% of administered therapeutics. Knowledge of how CYP3A4 adjusts and reshapes the active site to regioselectively oxidize chemically diverse compounds is critical for better understanding structure-function relations in this important enzyme, improving the outcomes for drug metabolism predictions, and developing pharmaceuticals that have a decreased ability to undergo metabolism and cause detrimental drug-drug interactions. However, there is very limited structural information on CYP3A4-substrate interactions available to date. Despite the vast variety of drugs undergoing metabolism, only the sedative midazolam (MDZ) serves as a marker substrate for the in vivo activity assessment because it is preferentially and regioselectively oxidized by CYP3A4. We solved the 2.7 Å crystal structure of the CYP3A4-MDZ complex, where the drug is well defined and oriented suitably for hydroxylation of the C1 atom, the major site of metabolism. This binding mode requires H-bonding to Ser119 and a dramatic conformational switch in the F-G fragment, which transmits to the adjacent D, E, H, and I helices, resulting in a collapse of the active site cavity and MDZ immobilization. In addition to providing insights on the substrate-triggered active site reshaping (an induced fit), the crystal structure explains the accumulated experimental results, identifies possible effector binding sites, and suggests why MDZ is predominantly metabolized by the CYP3A enzyme subfamily.
Organizational Affiliation:
Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697-3900; [email protected].