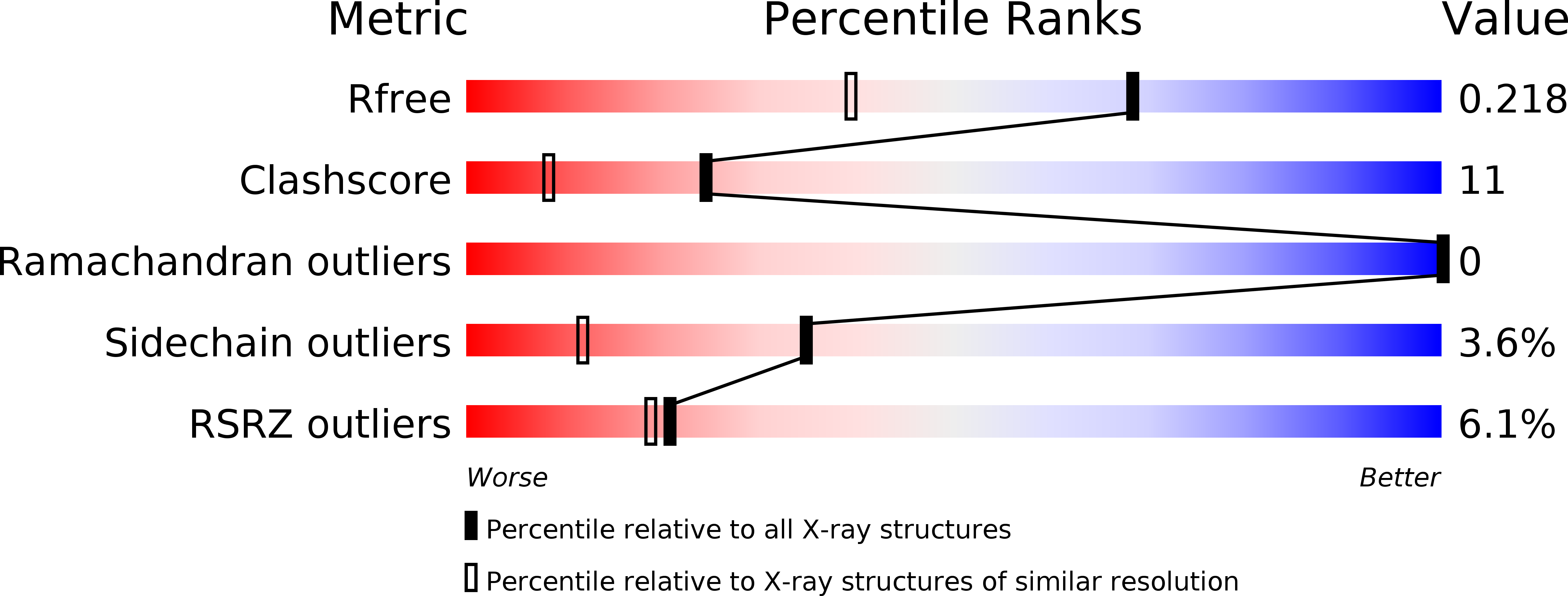
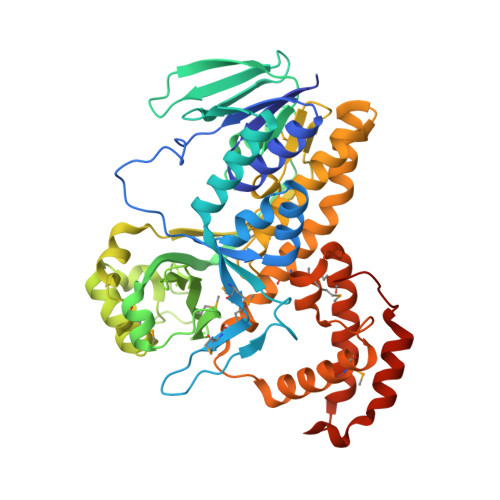
Biochemistry and structural studies of kynurenine 3-monooxygenase reveal allosteric inhibition by Ro 61-8048
Gao, J., Yao, L., Xia, T., Liao, X., Zhu, D., Xiang, Y.(2018) FASEB J 32: 2036-2045
- PubMed: 29208702
- DOI: https://doi.org/10.1096/fj.201700397RR
- Primary Citation of Related Structures:
5Y66, 5Y77, 5Y7A - PubMed Abstract:
The human kynurenine 3-monooxygenase (hKMO) is a potential therapeutic target for neurodegenerative and neurologic disorders. Inhibition of KMO by Ro 61-8048, a potent, selective, and the most widely used inhibitor of KMO, was shown effective in various models of neurodegenerative or neurologic disorders. However, the molecular basis of hKMO inhibition by Ro 61-8048 is not clearly understood. Here, we report biochemistry studies on hKMO and crystal structures of an hKMO homolog, pfKMO from Pseudomonas fluorescens, in complex with the substrate l-kynurenine and Ro 61-8048. We found that the C-terminal ∼110 aa are essential for the enzymatic activity of hKMO and the homologous C-terminal region of pfKMO folds into a distinct, all-α-helical domain, which associates with the N-terminal catalytic domain to form a unique tunnel in proximity to the substrate-binding pocket. The tunnel binds the Ro 61-8048 molecule, which fills most of the tunnel, and Ro 61-8048 is hydrogen bonded with several completely conserved residues, including an essential catalytic residue. Modification of Ro 61-8048 and biochemical studies of the modified Ro 61-8048 derivatives suggested that Ro 61-8048 inhibits the enzyme in an allosteric manner by affecting the conformation of the essential catalytic residue and by blocking entry of the substrate or product release. The unique binding sites distinguish Ro 61-8048 as a noncompetitive and highly selective inhibitor from other competitive inhibitors, which should facilitate further optimization of Ro 61-8048 and the development of new inhibitory drugs to hKMO.-Gao, J., Yao, L., Xia, T., Liao, X., Zhu, D., Xiang, Y. Biochemistry and structural studies of kynurenine 3-monooxygenase reveal allosteric inhibition by Ro 61-8048.
Organizational Affiliation:
School of Medicine, Center for Infectious Disease Research, Collaborative Innovation Center for Diagnosis and Treatment of Infectious Diseases, Beijing Advanced Innovation Center for Structural Biology.