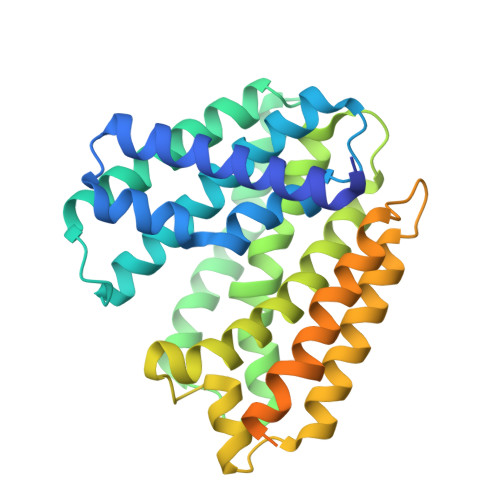
Moenomycin Biosynthesis: Structure and Mechanism of Action of the Prenyltransferase MoeN5.
Zhang, L., Chen, C.C., Ko, T.P., Huang, J.W., Zheng, Y., Liu, W., Wang, I., Malwal, S.R., Feng, X., Wang, K., Huang, C.H., Hsu, S.T., Wang, A.H., Oldfield, E., Guo, R.T.(2016) Angew Chem Int Ed Engl 55: 4716-4720
- PubMed: 26954060
- DOI: https://doi.org/10.1002/anie.201511388
- Primary Citation of Related Structures:
5B00, 5B02, 5B03, 5B0I, 5B0J, 5B0K, 5B0L, 5B0M - PubMed Abstract:
The structure of MoeN5, a unique prenyltransferase involved in the biosynthesis of the antibiotic moenomycin, is reported. MoeN5 catalyzes the reaction of geranyl diphosphate (GPP) with the cis-farnesyl group in phosphoglycolipid 5 to form the (C25) moenocinyl-sidechain-containing lipid 7. GPP binds to an allylic site (S1) and aligns well with known S1 inhibitors. Alkyl glycosides, glycolipids, can bind to both S1 and a second site, S2. Long sidechains in S2 are "bent" and co-locate with the homoallylic substrate isopentenyl diphosphate in other prenyltransferases. These observations support a MoeN5 mechanism in which 5 binds to S2 with its C6-C11 group poised to attack C1 in GPP to form the moenocinyl sidechain, with the more distal regions of 5 aligning with the distal glucose in decyl maltoside. The results are of general interest because they provide the first structures of MoeN5 and a structural basis for its mechanism of action, results that will facilitate the design of new antibiotics.
Organizational Affiliation:
Industrial Enzymes National Engineering Laboratory, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin, 300308, China.