HIV-1 CA hexamer with NUP153 peptide - R3 crystal form
Skorupka, K., Zadrozny, K., Ganser-Pornillos, B.K., Pornillos, O.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| HIV-1 CA protein | 231 | Human immunodeficiency virus type 1 (NEW YORK-5 ISOLATE) | Mutation(s): 4 Gene Names: gag | 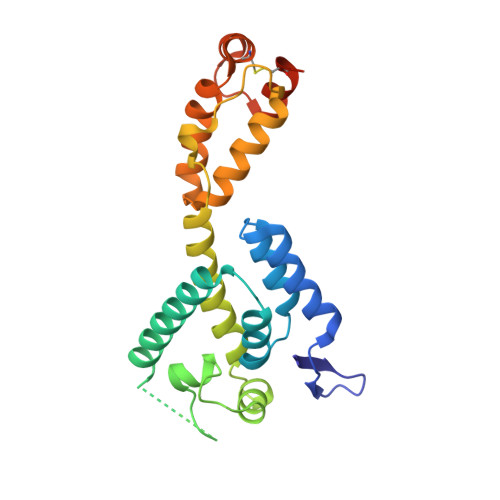 | |
UniProt | |||||
Find proteins for P12493 (Human immunodeficiency virus type 1 group M subtype B (isolate NY5)) Explore P12493 Go to UniProtKB: P12493 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P12493 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Nuclear pore complex protein Nup153 | C [auth D] | 23 | Homo sapiens | Mutation(s): 0 |  |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P49790 (Homo sapiens) Explore P49790 Go to UniProtKB: P49790 | |||||
PHAROS: P49790 GTEx: ENSG00000124789 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P49790 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| FLU Query on FLU | D [auth B] | 2-(6-HYDROXY-3-OXO-3H-XANTHEN-9-YL)-BENZOIC ACID C20 H12 O5 YKGGGCXBWXHKIZ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 151.924 | α = 90 |
| b = 151.924 | β = 90 |
| c = 69.646 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | R01-AI120956 |
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | P50-GM103297 |