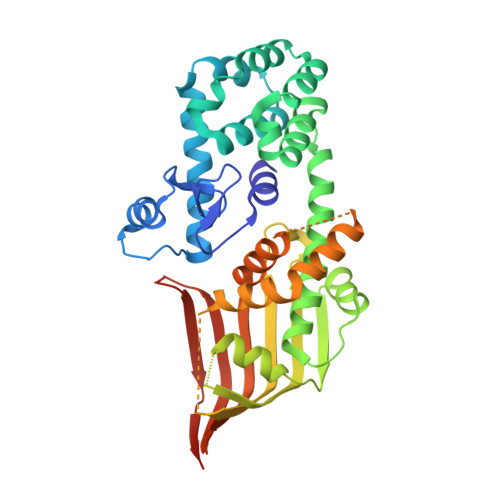
Atg2 mediates direct lipid transfer between membranes for autophagosome formation.
Osawa, T., Kotani, T., Kawaoka, T., Hirata, E., Suzuki, K., Nakatogawa, H., Ohsumi, Y., Noda, N.N.(2019) Nat Struct Mol Biol 26: 281-288
- PubMed: 30911189
- DOI: https://doi.org/10.1038/s41594-019-0203-4
- Primary Citation of Related Structures:
6A9E, 6A9J - PubMed Abstract:
A key event in autophagy is autophagosome formation, whereby the newly synthesized isolation membrane (IM) expands to form a complete autophagosome using endomembrane-derived lipids. Atg2 physically links the edge of the expanding IM with the endoplasmic reticulum (ER), a role that is essential for autophagosome formation. However, the molecular function of Atg2 during ER-IM contact remains unclear, as does the mechanism of lipid delivery to the IM. Here we show that the conserved amino-terminal region of Schizosaccharomyces pombe Atg2 includes a lipid-transfer-protein-like hydrophobic cavity that accommodates phospholipid acyl chains. Atg2 bridges highly curved liposomes, thereby facilitating efficient phospholipid transfer in vitro, a function that is inhibited by mutations that impair autophagosome formation in vivo. These results suggest that Atg2 acts as a lipid-transfer protein that supplies phospholipids for autophagosome formation.
Organizational Affiliation:
Institute of Microbial Chemistry (BIKAKEN), Tokyo, Japan.