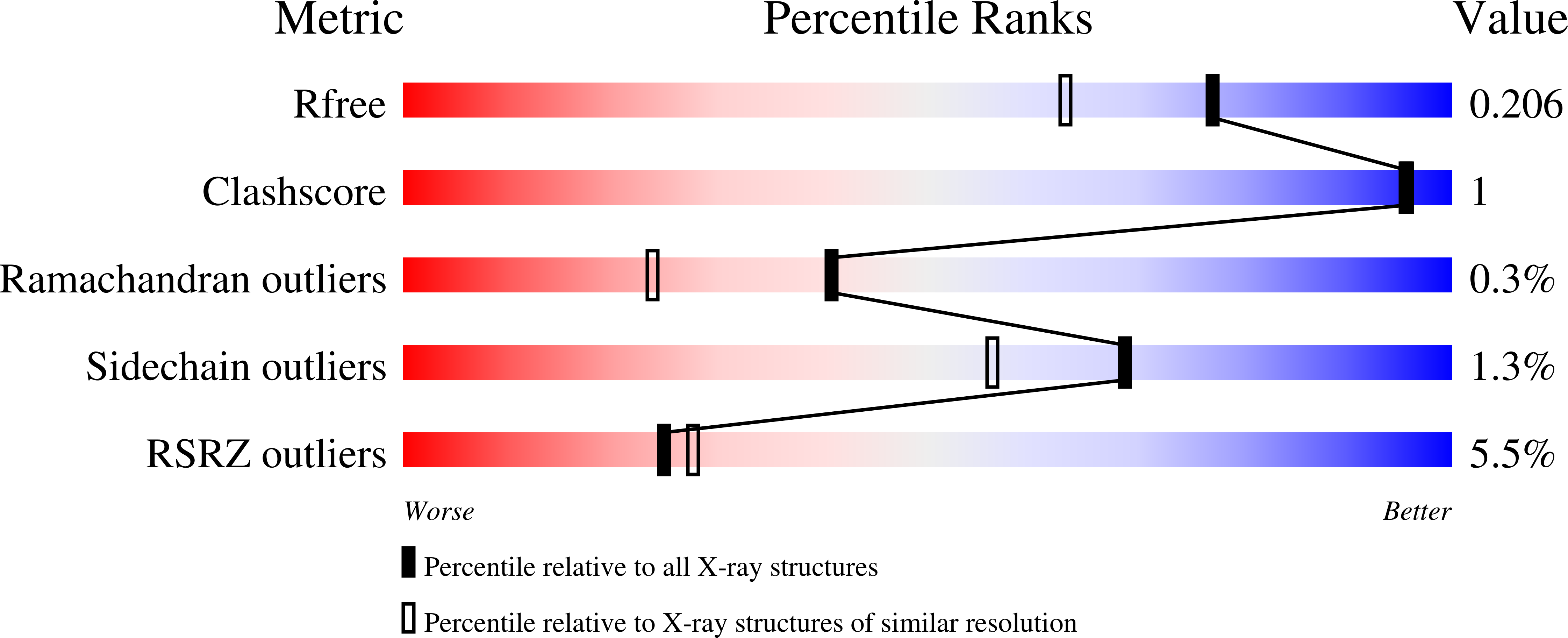
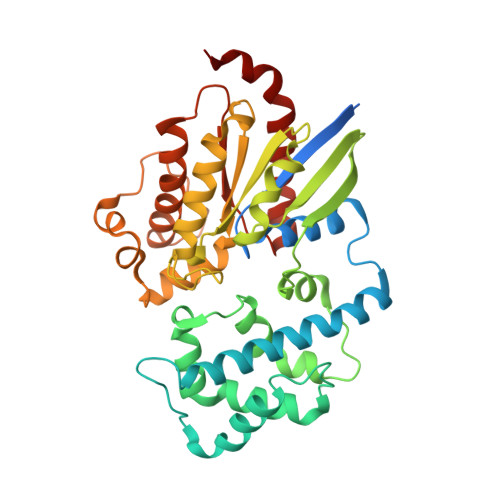
Disease-Causing Mutations in the G Protein G alpha s Subvert the Roles of GDP and GTP.
Hu, Q., Shokat, K.M.(2018) Cell 173: 1254-1264.e11
- PubMed: 29628140
- DOI: https://doi.org/10.1016/j.cell.2018.03.018
- Primary Citation of Related Structures:
6AU6 - PubMed Abstract:
The single most frequent cancer-causing mutation across all heterotrimeric G proteins is R201C in Gαs. The current model explaining the gain-of-function activity of the R201 mutations is through the loss of GTPase activity and resulting inability to switch off to the GDP state. Here, we find that the R201C mutation can bypass the need for GTP binding by directly activating GDP-bound Gαs through stabilization of an intramolecular hydrogen bond network. Having found that a gain-of-function mutation can convert GDP into an activator, we postulated that a reciprocal mutation might disrupt the normal role of GTP. Indeed, we found R228C, a loss-of-function mutation in Gαs that causes pseudohypoparathyroidism type 1a (PHP-Ia), compromised the adenylyl cyclase-activating activity of Gαs bound to a non-hydrolyzable GTP analog. These findings show that disease-causing mutations in Gαs can subvert the canonical roles of GDP and GTP, providing new insights into the regulation mechanism of G proteins.
Organizational Affiliation:
Department of Cellular and Molecular Pharmacology and Howard Hughes Medical Institute, University of California-San Francisco, San Francisco, CA 94158, USA.