hPDE4D2 structure with inhibitor NPD-1439
Salado, I.G., Moreno, C., Sakaine, G., Singh, A.K., Blaazer, A.R., Siderius, M., Matheeussen, A., Gul, S., Maes, L., Leurs, R., Brown, D.G., Augustyns, K.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cAMP-specific 3',5'-cyclic phosphodiesterase 4D | 364 | Homo sapiens | Mutation(s): 0 Gene Names: PDE4D, DPDE3 EC: 3.1.4.53 | 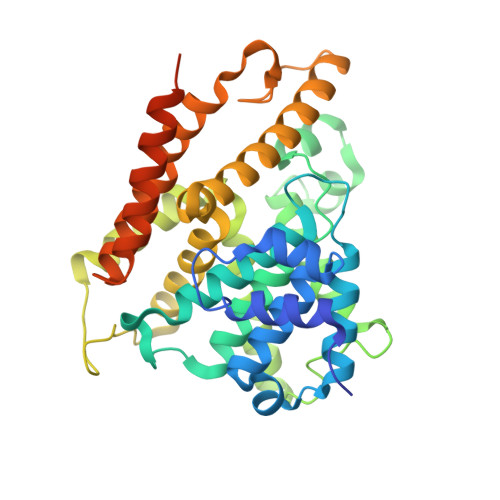 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q08499 (Homo sapiens) Explore Q08499 Go to UniProtKB: Q08499 | |||||
PHAROS: Q08499 GTEx: ENSG00000113448 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q08499 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 8 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| D68 (Subject of Investigation/LOI) Query on D68 | KC [auth D], NB [auth C], QA [auth B], U [auth A] | 1-(2-{4-[(4aS,8aR)-4-[3,4-bis(difluoromethoxy)phenyl]-1-oxo-1,2,4a,5,8,8a-hexahydrophthalazin-2-yl]piperidin-1-yl}-2-oxoethyl)-4,4-dimethylpiperidine-2,6-dione C30 H34 F4 N4 O6 WVAQTVIALKSAMT-VQTJNVASSA-N |  | ||
| EPE Query on EPE | JA [auth B], KB [auth C], LC [auth D], PA [auth B], V [auth A] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| PG4 Query on PG4 | MB [auth C], SB [auth D] | TETRAETHYLENE GLYCOL C8 H18 O5 UWHCKJMYHZGTIT-UHFFFAOYSA-N |  | ||
| PEG Query on PEG | JB [auth C], JC [auth D], N [auth A], OA [auth B] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| ZN Query on ZN | CB [auth C], DA [auth B], E [auth A], UB [auth D] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | AA [auth A] AB [auth C] AC [auth D] BA [auth A] BB [auth C] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| NI Query on NI | CA [auth A], RC [auth D] | NICKEL (II) ION Ni VEQPNABPJHWNSG-UHFFFAOYSA-N |  | ||
| MG Query on MG | DB [auth C], EA [auth B], F [auth A], VB [auth D] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 97.19 | α = 90 |
| b = 110.71 | β = 90 |
| c = 161.48 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| REFMAC | phasing |