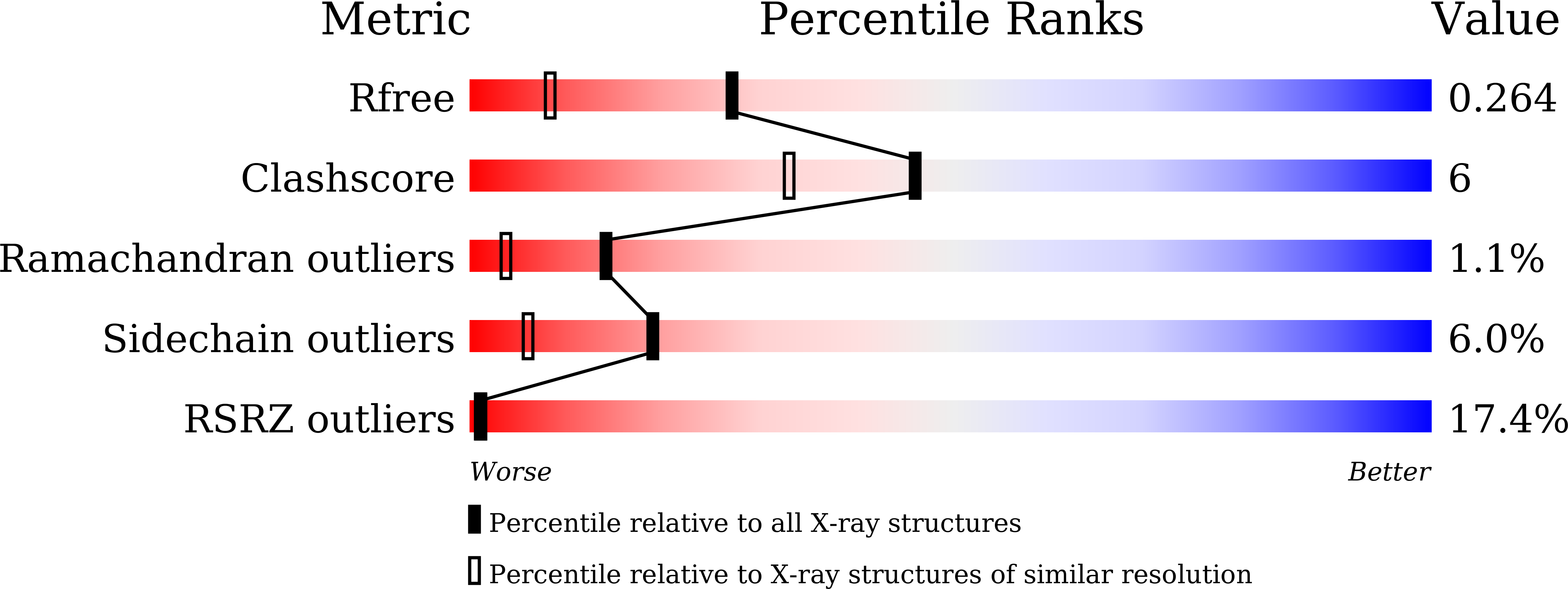
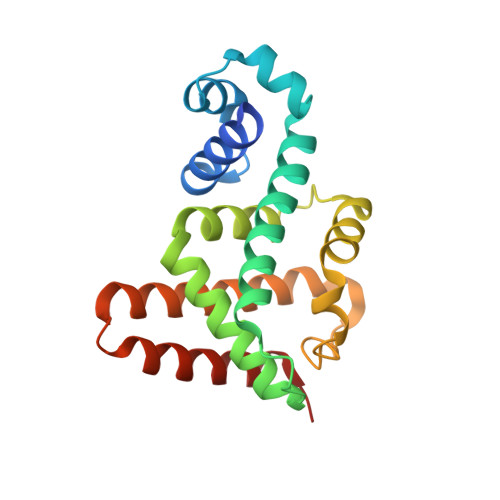
Crystal structure of the multidrug resistance regulator RamR complexed with bile acids.
Yamasaki, S., Nakashima, R., Sakurai, K., Baucheron, S., Giraud, E., Doublet, B., Cloeckaert, A., Nishino, K.(2019) Sci Rep 9: 177-177
- PubMed: 30655545
- DOI: https://doi.org/10.1038/s41598-018-36025-8
- Primary Citation of Related Structures:
6IE8, 6IE9 - PubMed Abstract:
During infection, Salmonella senses and responds to harsh environments within the host. Persistence in a bile-rich environment is important for Salmonella to infect the small intestine or gallbladder and the multidrug efflux system AcrAB-TolC is required for bile resistance. The genes encoding this system are mainly regulated by the ramRA locus, which is composed of the divergently transcribed ramA and ramR genes. The acrAB and tolC genes are transcriptionally activated by RamA, whose encoding gene is itself transcriptionally repressed by RamR. RamR recognizes multiple drugs; however, the identity of the environmental signals to which it responds is unclear. Here, we describe the crystal structures of RamR in complexes with bile components, including cholic acid and chenodeoxycholic acid, determined at resolutions of 2.0 and 1.8 Å, respectively. Both cholic and chenodeoxycholic acids form four hydrogen bonds with Tyr59, Thr85, Ser137 and Asp152 of RamR, instead of π-π interactions with Phe155, a residue that is important for the recognition of multiple compounds including berberine, crystal violet, dequalinium, ethidium bromide and rhodamine 6 G. Binding of these compounds to RamR reduces its DNA-binding affinity, resulting in the increased transcription of ramA and acrAB-tolC. Our results reveal that Salmonella senses bile acid components through RamR and then upregulates the expression of RamA, which can lead to induction of acrAB-tolC expression with resulting tolerance to bile-rich environments.
Organizational Affiliation:
Graduate School of Pharmaceutical Sciences, Osaka University, 1-6 Yamadaoka, Suita, 565-0871, Osaka, Japan.