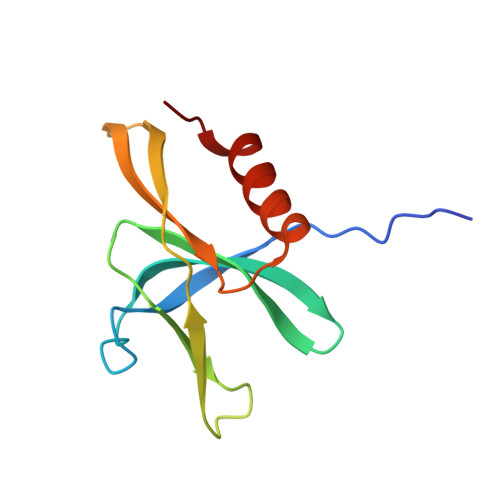
Crystal Structure of the SARS-CoV-2 Non-structural Protein 9, Nsp9.
Littler, D.R., Gully, B.S., Colson, R.N., Rossjohn, J.(2020) iScience 23: 101258-101258
- PubMed: 32592996
- DOI: https://doi.org/10.1016/j.isci.2020.101258
- Primary Citation of Related Structures:
6W9Q, 6WXD - PubMed Abstract:
Many of the SARS-CoV-2 proteins have related counterparts across the Severe Acute Respiratory Syndrome (SARS-CoV) family. One such protein is non-structural protein 9 (Nsp9), which is thought to mediate viral replication, overall virulence, and viral genomic RNA reproduction. We sought to better characterize the SARS-CoV-2 Nsp9 and subsequently solved its X-ray crystal structure, in an apo form and, unexpectedly, in a peptide-bound form with a sequence originating from a rhinoviral 3C protease sequence (LEVL). The SARS-CoV-2 Nsp9 structure revealed the high level of structural conservation within the Nsp9 family. The exogenous peptide binding site is close to the dimer interface and impacted the relative juxtapositioning of the monomers within the homodimer. We have established a protocol for the production of SARS-CoV-2 Nsp9, determined its structure, and identified a peptide-binding site that warrants further study to understanding Nsp9 function.
Organizational Affiliation:
Infection and Immunity Program, Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, VIC, Australia; Australian Research Council Centre of Excellence for Advanced Molecular Imaging, Monash University, Clayton, VIC, Australia. Electronic address: [email protected].