Tight and specific lanthanide binding in a de novo TIM barrel with a large internal cavity designed by symmetric domain fusion.
Caldwell, S.J., Haydon, I.C., Piperidou, N., Huang, P.S., Bick, M.J., Sjostrom, H.S., Hilvert, D., Baker, D., Zeymer, C.(2020) Proc Natl Acad Sci U S A 117: 30362-30369
- PubMed: 33203677
- DOI: https://doi.org/10.1073/pnas.2008535117
- Primary Citation of Related Structures:
6WVS, 6WXO, 6WXP, 6ZV9 - PubMed Abstract:
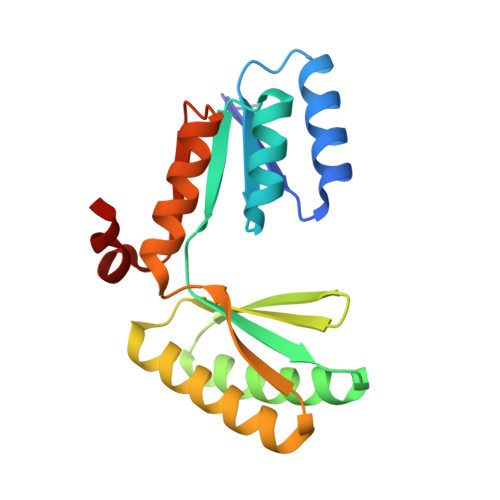
De novo protein design has succeeded in generating a large variety of globular proteins, but the construction of protein scaffolds with cavities that could accommodate large signaling molecules, cofactors, and substrates remains an outstanding challenge. The long, often flexible loops that form such cavities in many natural proteins are difficult to precisely program and thus challenging for computational protein design. Here we describe an alternative approach to this problem. We fused two stable proteins with C2 symmetry-a de novo designed dimeric ferredoxin fold and a de novo designed TIM barrel-such that their symmetry axes are aligned to create scaffolds with large cavities that can serve as binding pockets or enzymatic reaction chambers. The crystal structures of two such designs confirm the presence of a 420 cubic Ångström chamber defined by the top of the designed TIM barrel and the bottom of the ferredoxin dimer. We functionalized the scaffold by installing a metal-binding site consisting of four glutamate residues close to the symmetry axis. The protein binds lanthanide ions with very high affinity as demonstrated by tryptophan-enhanced terbium luminescence. This approach can be extended to other metals and cofactors, making this scaffold a modular platform for the design of binding proteins and biocatalysts.
Organizational Affiliation:
Department of Biochemistry, University of Washington, Seattle, WA 98195.