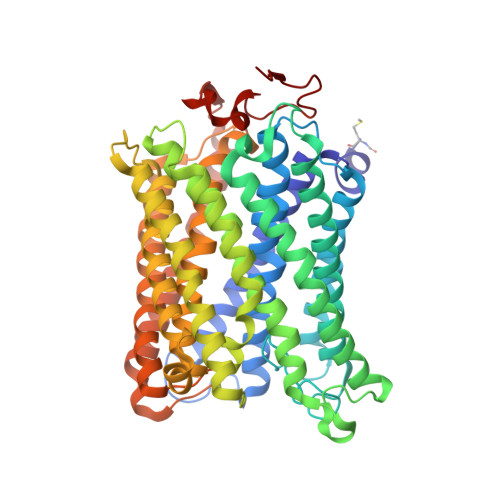
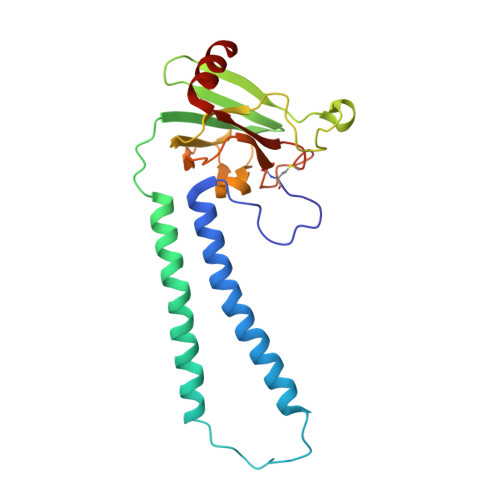
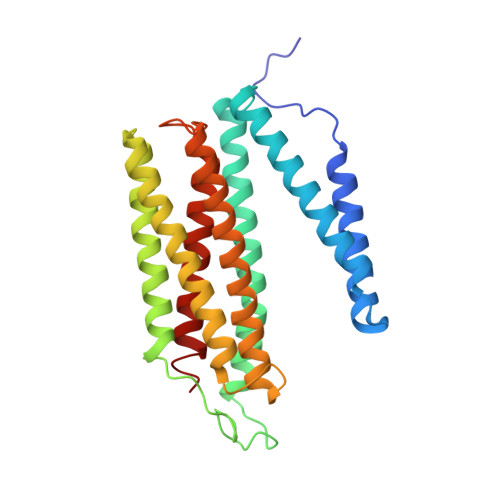
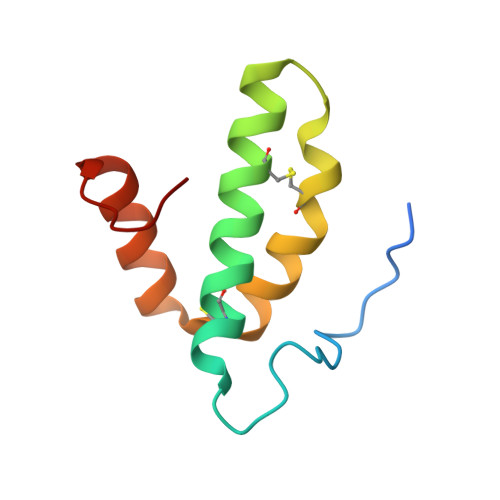
Snapshot of an oxygen intermediate in the catalytic reaction of cytochromecoxidase.
Ishigami, I., Lewis-Ballester, A., Echelmeier, A., Brehm, G., Zatsepin, N.A., Grant, T.D., Coe, J.D., Lisova, S., Nelson, G., Zhang, S., Dobson, Z.F., Boutet, S., Sierra, R.G., Batyuk, A., Fromme, P., Fromme, R., Spence, J.C.H., Ros, A., Yeh, S.R., Rousseau, D.L.(2019) Proc Natl Acad Sci U S A 116: 3572-3577
- PubMed: 30808749
- DOI: https://doi.org/10.1073/pnas.1814526116
- Primary Citation of Related Structures:
6NKN, 6NMF, 6NMP - PubMed Abstract:
Cytochrome c oxidase (C c O) reduces dioxygen to water and harnesses the chemical energy to drive proton translocation across the inner mitochondrial membrane by an unresolved mechanism. By using time-resolved serial femtosecond crystallography, we identified a key oxygen intermediate of bovine C c O. It is assigned to the P R -intermediate, which is characterized by specific redox states of the metal centers and a distinct protein conformation. The heme a 3 iron atom is in a ferryl (Fe 4+ = O 2- ) configuration, and heme a and Cu B are oxidized while Cu A is reduced. A Helix-X segment is poised in an open conformational state; the heme a farnesyl sidechain is H-bonded to S382, and loop-I-II adopts a distinct structure. These data offer insights into the mechanism by which the oxygen chemistry is coupled to unidirectional proton translocation.
Organizational Affiliation:
Department of Physiology and Biophysics, Albert Einstein College of Medicine, Bronx, NY 10461.