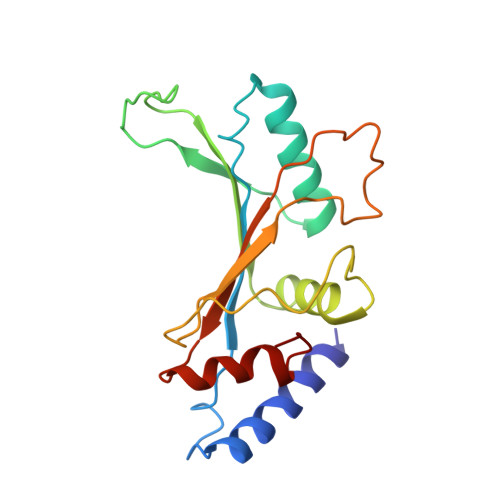
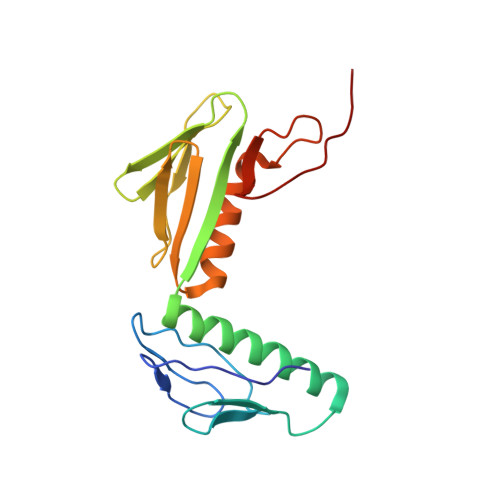
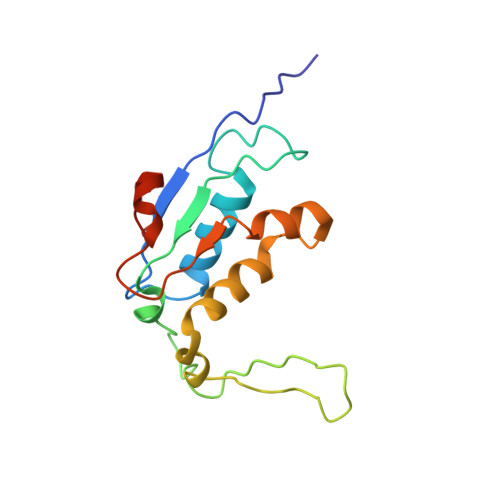
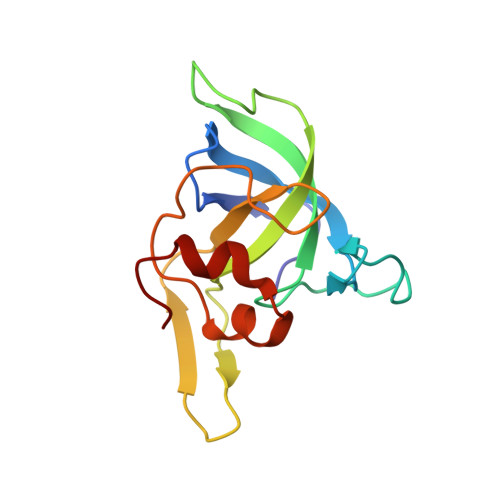
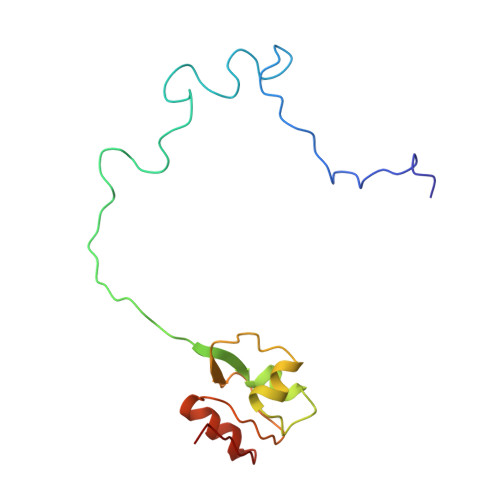
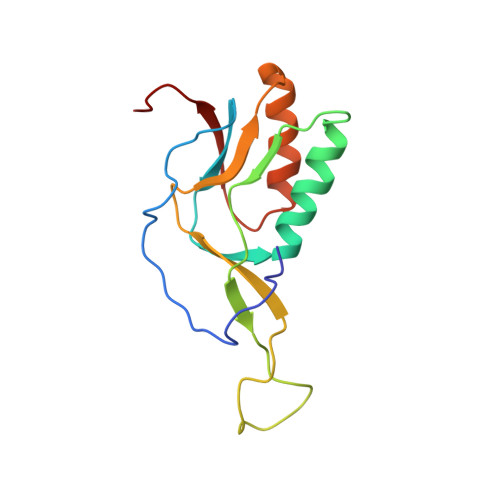
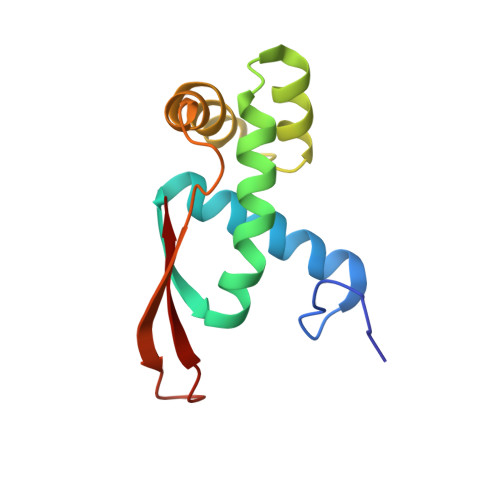
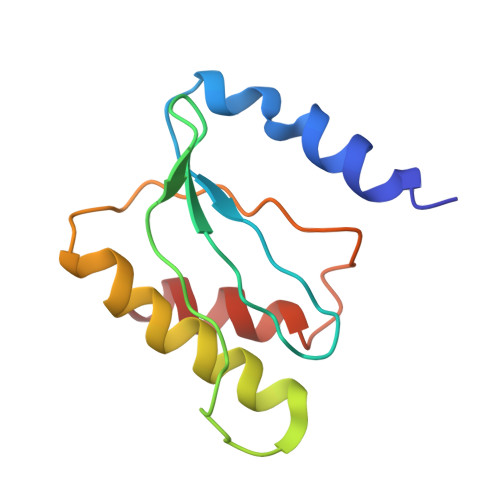
Ribosome-binding and anti-microbial studies of the mycinamicins, 16-membered macrolide antibiotics from Micromonospora griseorubida.
Breiner-Goldstein, E., Eyal, Z., Matzov, D., Halfon, Y., Cimicata, G., Baum, M., Rokney, A., Ezernitchi, A.V., Lowell, A.N., Schmidt, J.J., Rozenberg, H., Zimmerman, E., Bashan, A., Valinsky, L., Anzai, Y., Sherman, D.H., Yonath, A.(2021) Nucleic Acids Res 49: 9560-9573
- PubMed: 34417608
- DOI: https://doi.org/10.1093/nar/gkab684
- Primary Citation of Related Structures:
7A0R, 7A0S, 7A18 - PubMed Abstract:
Macrolides have been effective clinical antibiotics for over 70 years. They inhibit protein biosynthesis in bacterial pathogens by narrowing the nascent protein exit tunnel in the ribosome. The macrolide class of natural products consist of a macrolactone ring linked to one or more sugar molecules. Most of the macrolides used currently are semi-synthetic erythromycin derivatives, composed of a 14- or 15-membered macrolactone ring. Rapidly emerging resistance in bacterial pathogens is among the most urgent global health challenges, which render many antibiotics ineffective, including next-generation macrolides. To address this threat and advance a longer-term plan for developing new antibiotics, we demonstrate how 16-membered macrolides overcome erythromycin resistance in clinically isolated Staphylococcus aureus strains. By determining the structures of complexes of the large ribosomal subunit of Deinococcus radiodurans (D50S) with these 16-membered selected macrolides, and performing anti-microbial studies, we identified resistance mechanisms they may overcome. This new information provides important insights toward the rational design of therapeutics that are effective against drug resistant human pathogens.
Organizational Affiliation:
Department of Chemical and Structural Biology, The Weizmann Institute of Science, Rehovot 760001, Israel.