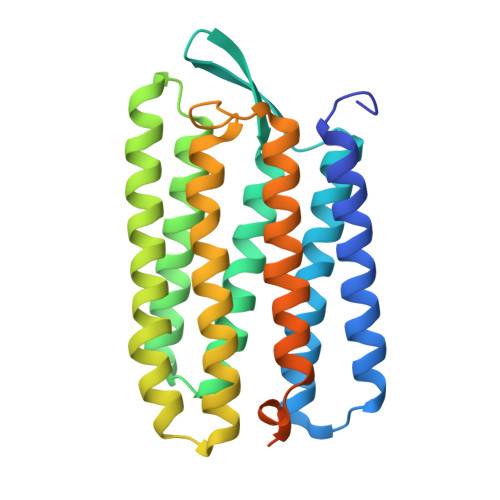
High-pressure crystallography shows noble gas intervention into protein-lipid interaction and suggests a model for anaesthetic action.
Melnikov, I., Orekhov, P., Rulev, M., Kovalev, K., Astashkin, R., Bratanov, D., Ryzhykau, Y., Balandin, T., Bukhdruker, S., Okhrimenko, I., Borshchevskiy, V., Bourenkov, G., Mueller-Dieckmann, C., van der Linden, P., Carpentier, P., Leonard, G., Gordeliy, V., Popov, A.(2022) Commun Biol 5: 360-360
- PubMed: 35422073
- DOI: https://doi.org/10.1038/s42003-022-03233-y
- Primary Citation of Related Structures:
7Q35, 7Q36, 7Q37, 7Q38 - PubMed Abstract:
In this work we examine how small hydrophobic molecules such as inert gases interact with membrane proteins (MPs) at a molecular level. High pressure atmospheres of argon and krypton were used to produce noble gas derivatives of crystals of three well studied MPs (two different proton pumps and a sodium light-driven ion pump). The structures obtained using X-ray crystallography showed that the vast majority of argon and krypton binding sites were located on the outer hydrophobic surface of the MPs - a surface usually accommodating hydrophobic chains of annular lipids (which are known structural and functional determinants for MPs). In conformity with these results, supplementary in silico molecular dynamics (MD) analysis predicted even greater numbers of argon and krypton binding positions on MP surface within the bilayer. These results indicate a potential importance of such interactions, particularly as related to the phenomenon of noble gas-induced anaesthesia.
Organizational Affiliation:
Institute of Biological Information Processing (IBI-7: Structural Biochemistry), Forschungszentrum Jülich GmbH, Jülich, Germany.