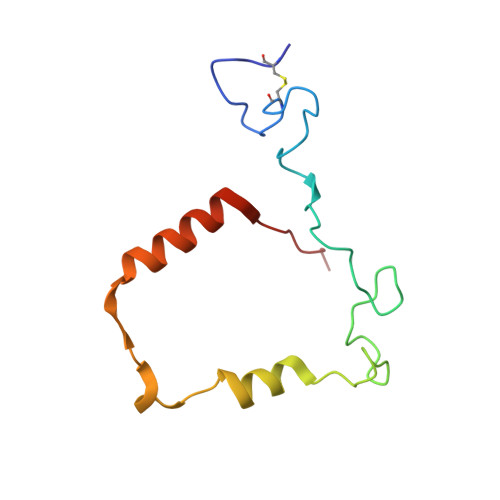
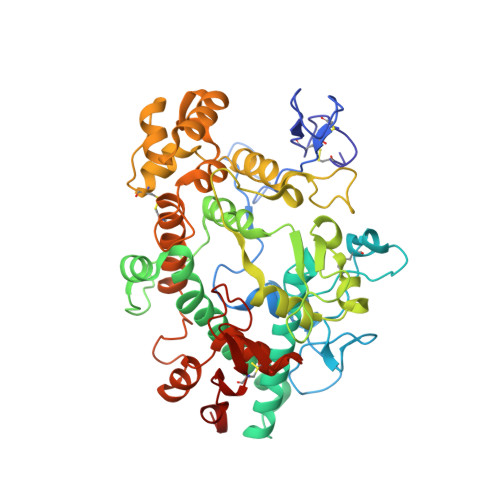
Posttranslational modification and heme cavity architecture of human eosinophil peroxidase-insights from first crystal structure and biochemical characterization.
Pfanzagl, V., Gruber-Grunwald, C., Leitgeb, U., Furtmuller, P.G., Obinger, C.(2023) J Biol Chem 299: 105402-105402
- PubMed: 38229400
- DOI: https://doi.org/10.1016/j.jbc.2023.105402
- Primary Citation of Related Structures:
8OGI - PubMed Abstract:
Eosinophil peroxidase (EPO) is the most abundant granule protein exocytosed by eosinophils, specialized human phagocytes. Released EPO catalyzes the formation of reactive oxidants from bromide, thiocyanate, and nitrite that kill tissue-invading parasites. However, EPO also plays a deleterious role in inflammatory diseases, making it a potential pharmacological target. A major hurdle is the high similarity to the homologous myeloperoxidase (MPO), which requires a detailed understanding of the small structural differences that can be used to increase the specificity of the inhibitors. Here, we present the first crystal structure of mature leukocyte EPO at 1.6 Å resolution together with analyses of its posttranslational modifications and biochemical properties. EPO has an exceptionally high number of positively charged surface patches but only two occupied glycosylation sites. The crystal structure further revealed the existence of a light (L) and heavy (H) chain as a result of proteolytic cleavage. Detailed comparison with the structure of human MPO allows us to identify differences that may contribute to the known divergent enzymatic properties. The crystal structure revealed fully established ester links between the prosthetic group and the protein, the comparably weak imidazolate character of the proximal histidine, and the conserved structure of the catalytic amino acids and Ca 2+ -binding site. Prediction of the structure of unprocessed proeosinophil peroxidase allows further structural analysis of the three protease cleavage sites and the potential pro-convertase recognition site in the propeptide. Finally, EPO biosynthesis and its biochemical and biophysical properties are discussed with respect to the available data from the well-studied MPO.
Organizational Affiliation:
Department of Chemistry, Institute of Biochemistry, University of Natural Resources and Life Sciences, Vienna, Austria. Electronic address: [email protected].