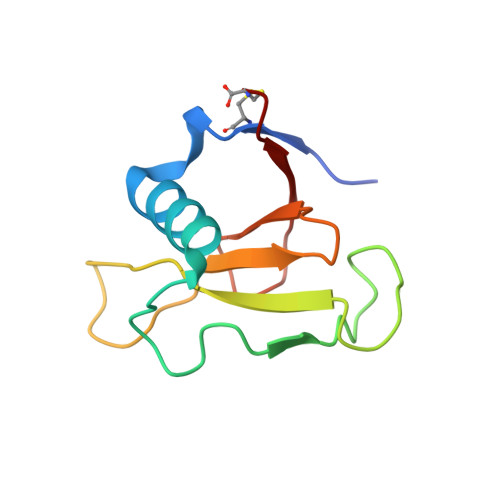
Crystal structure reveals two alternative conformations in the active site of ribonuclease Sa2.
Sevcik, J., Dauter, Z., Wilson, K.S.(2004) Acta Crystallogr D Biol Crystallogr 60: 1198-1204
- PubMed: 15213380
- DOI: https://doi.org/10.1107/S0907444904009035
- Primary Citation of Related Structures:
1PY3, 1PYL - PubMed Abstract:
Three different strains of Streptomyces aureofaciens produce the homologous ribonucleases Sa, Sa2 and Sa3. The crystal structures of ribonuclease Sa (RNase Sa) and its complexes with mononucleotides have previously been reported at high resolution. Here, the structures of two crystal forms (I and II) of ribonuclease Sa2 (RNase Sa2) are presented at 1.8 and 1.5 A resolution. The structures were determined by molecular replacement using the coordinates of RNase Sa as a search model and were refined to R factors of 17.5 and 15.0% and R(free) factors of 21.8 and 17.2%, respectively. The asymmetric unit of crystal form I contains three enzyme molecules, two of which have similar structures to those seen for ribonuclease Sa, with Tyr87 at the bottom of their active sites. In the third molecule, Tyr87 has moved substantially: the CA atom moves almost 5 A and the OH of the side chain moves 10 A, inserting itself into the active site of a neighbouring molecule at a similar position to that observed for the nucleotide base in RNase Sa complexes. The asymmetric unit of crystal form II contains two Sa2 molecules, both of which are similar to the usual Sa structures. In one molecule, two main-chain conformations were modelled in the alpha-helix. Finally, a brief comparison is made between the conformations of the Sa2 molecules and those of 34 independent molecules taken from 20 structures of ribonuclease Sa and two independent molecules taken from two structures of ribonuclease Sa3 in various crystal forms.
Organizational Affiliation:
Institute of Molecular Biology, Member of the Centre of Excellence for Molecular Medicine, Slovak Academy of Sciences, Dubravska Cesta 21, 84551 Bratislava, Slovak Republic. [email protected]