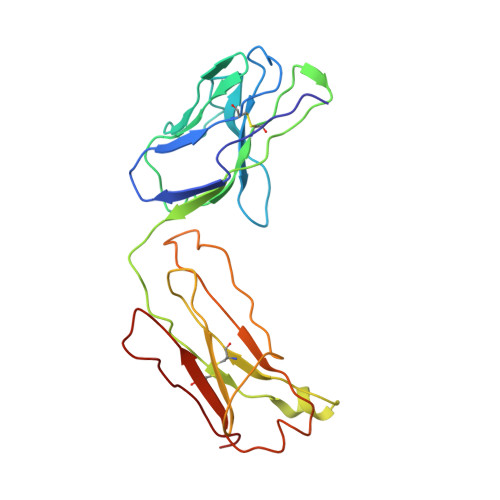
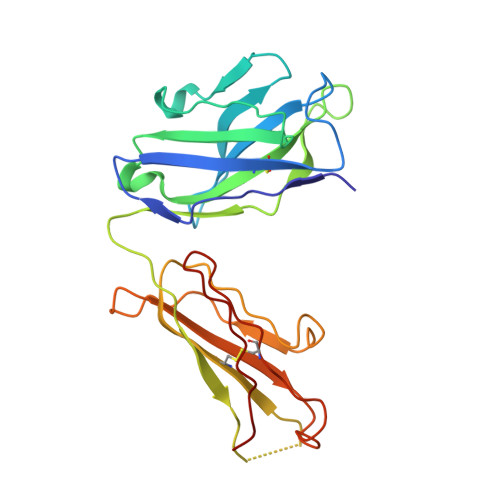
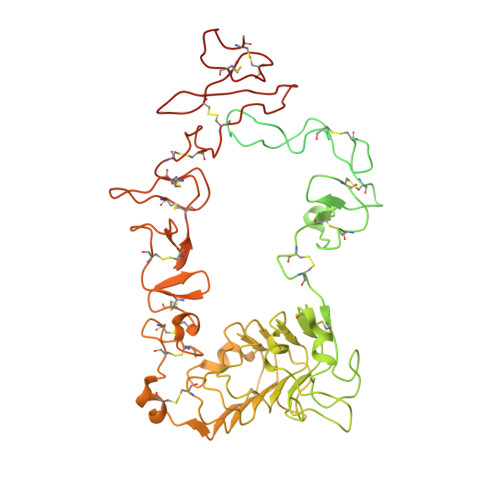
Structural basis for EGF receptor inhibition by the therapeutic antibody IMC-11F8.
Li, S., Kussie, P., Ferguson, K.M.(2008) Structure 16: 216-227
- PubMed: 18275813
- DOI: https://doi.org/10.1016/j.str.2007.11.009
- Primary Citation of Related Structures:
3B2U, 3B2V - PubMed Abstract:
Therapeutic anticancer strategies that target and inactivate the epidermal growth factor receptor (EGFR) are under intense study in the clinic. Here we describe the mechanism of EGFR inhibition by an antibody drug IMC-11F8. IMC-11F8 is a fully human antibody that has similar antitumor potency as the chimeric cetuximab/Erbitux and might represent a safer therapeutic alternative. We report the X-ray crystal structure of the Fab fragment of IMC-11F8 (Fab11F8) in complex with the entire extracellular region and with isolated domain III of EGFR. We compare this to our previous study of the cetuximab/EGFR interaction. Fab11F8 interacts with a remarkably similar epitope, but through a completely different set of interactions. Both the similarities and differences in binding of these two antibodies have important implications for the development of inhibitors that could exploit this same mechanism of EGFR inhibition.
Organizational Affiliation:
Department of Physiology, University of Pennsylvania School of Medicine, Philadelphia, PA 19104, USA.