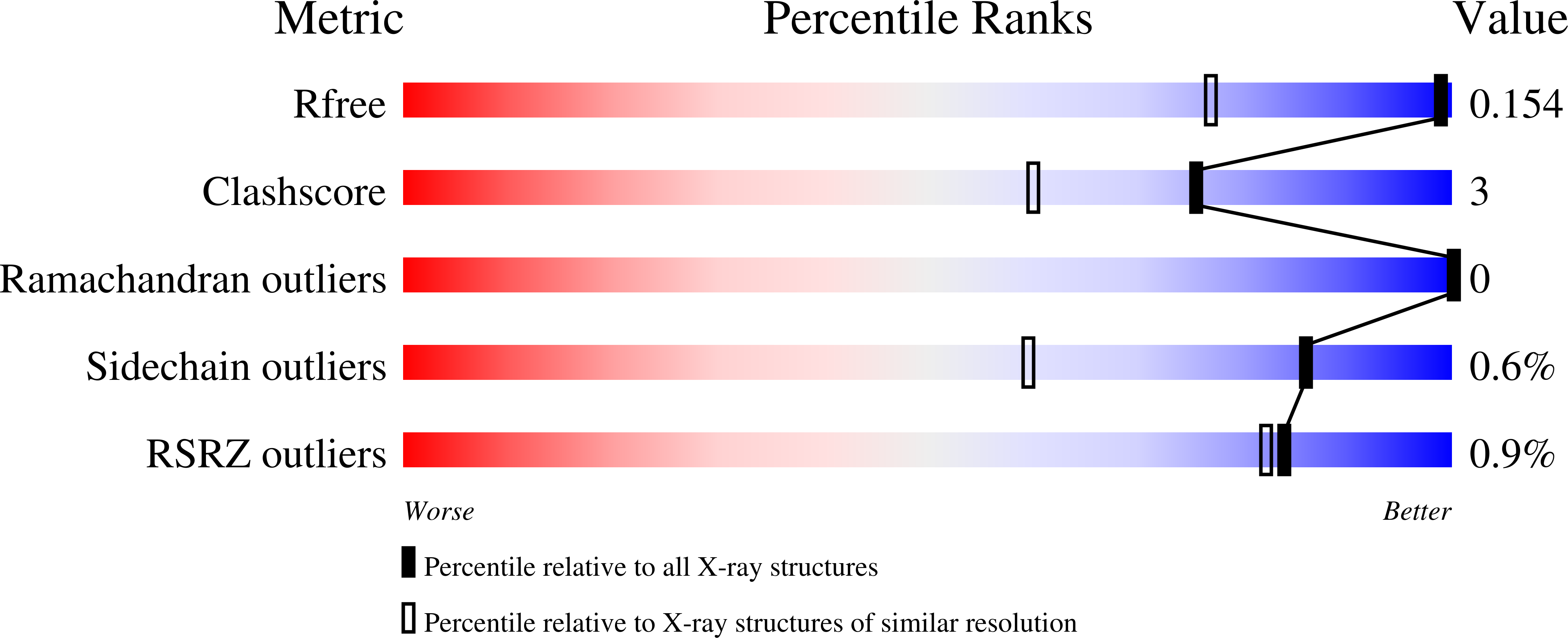
Crystallographic analyses of isoquinoline complexes reveal a new mode of metallo-beta-lactamase inhibition.
Li, G.B., Brem, J., Lesniak, R., Abboud, M.I., Lohans, C.T., Clifton, I.J., Yang, S.Y., Jimenez-Castellanos, J.C., Avison, M.B., Spencer, J., McDonough, M.A., Schofield, C.J.(2017) Chem Commun (Camb) 53: 5806-5809
- PubMed: 28470248
- DOI: https://doi.org/10.1039/c7cc02394d
- Primary Citation of Related Structures:
5N4S, 5N4T, 5N55, 5N58, 5NAI - PubMed Abstract:
Crystallographic analyses of the VIM-5 metallo-β-lactamase (MBL) with isoquinoline inhibitors reveal non zinc ion binding modes. Comparison with other MBL-inhibitor structures directed addition of a zinc-binding thiol enabling identification of potent B1 MBL inhibitors. The inhibitors potentiate meropenem activity against clinical isolates harboring MBLs.
Organizational Affiliation:
Department of Chemistry, University of Oxford, 12 Mansfield Road, Oxford, OX1 3TA, UK. [email protected] [email protected].