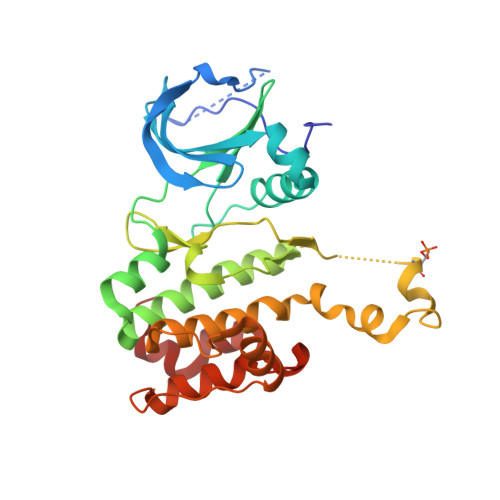
Redox priming promotes Aurora A activation during mitosis.
Lim, D.C., Joukov, V., Rettenmaier, T.J., Kumagai, A., Dunphy, W.G., Wells, J.A., Yaffe, M.B.(2020) Sci Signal 13
- PubMed: 32694171
- DOI: https://doi.org/10.1126/scisignal.abb6707
- Primary Citation of Related Structures:
6VPG, 6VPH, 6VPI, 6VPJ, 6VPL, 6VPM, 6XKA - PubMed Abstract:
Cell cycle-dependent redox changes can mediate transient covalent modifications of cysteine thiols to modulate the activities of regulatory kinases and phosphatases. Our previously reported finding that protein cysteine oxidation is increased during mitosis relative to other cell cycle phases suggests that redox modifications could play prominent roles in regulating mitotic processes. The Aurora family of kinases and their downstream targets are key components of the cellular machinery that ensures the proper execution of mitosis and the accurate segregation of chromosomes to daughter cells. In this study, x-ray crystal structures of the Aurora A kinase domain delineate redox-sensitive cysteine residues that, upon covalent modification, can allosterically regulate kinase activity and oligomerization state. We showed in both Xenopus laevis egg extracts and mammalian cells that a conserved cysteine residue within the Aurora A activation loop is crucial for Aurora A activation by autophosphorylation. We further showed that covalent disulfide adducts of this residue promote autophosphorylation of the Aurora A kinase domain. These findings reveal a potential mechanistic link between Aurora A activation and changes in the intracellular redox state during mitosis and provide insights into how novel small-molecule inhibitors may be developed to target specific subpopulations of Aurora A.
Organizational Affiliation:
MIT Center for Precision Cancer Medicine, Koch Institute for Integrative Cancer Research, and Departments of Biological Engineering and Biology, Massachusetts Institute of Technology, Cambridge, MA 02139, USA. [email protected] [email protected].