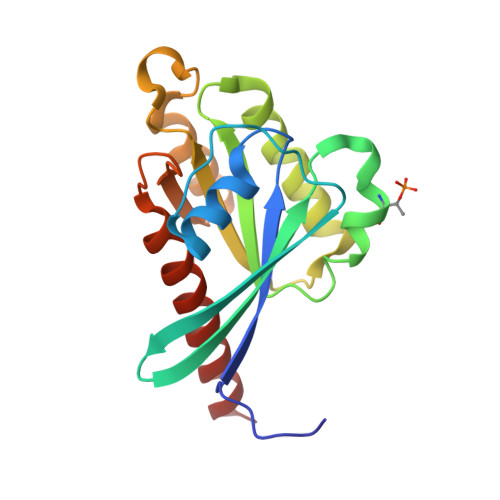
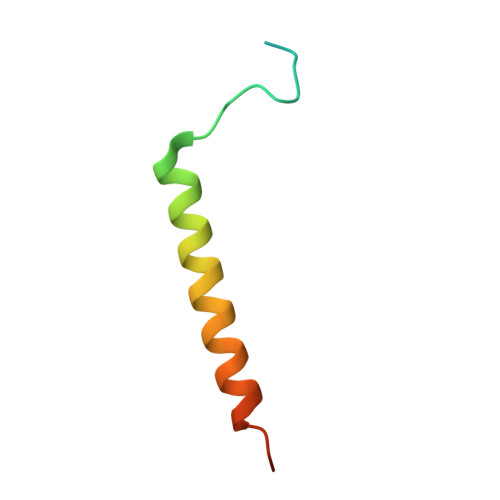
Dual arginine recognition of LRRK2 phosphorylated Rab GTPases.
Waschbusch, D., Purlyte, E., Khan, A.R.(2021) Biophys J 120: 1846-1855
- PubMed: 33887226
- DOI: https://doi.org/10.1016/j.bpj.2021.03.030
- Primary Citation of Related Structures:
6WHE, 7LWB - PubMed Abstract:
Parkinson's-disease-associated LRRK2 is a multidomain Ser/Thr kinase that phosphorylates a subset of Rab GTPases to control their effector functions. Rab GTPases are the prime regulators of membrane trafficking in eukaryotic cells. Rabs exert their biological effects by recruitment of effector proteins to subcellular compartments via their Rab-binding domain (RBD). Effectors are modular and typically contain additional domains that regulate various aspects of vesicle formation, trafficking, fusion, and organelle dynamics. The RBD of effectors is typically an α-helical coiled coil that recognizes the GTP conformation of the switch 1 and switch 2 motifs of Rabs. LRRK2 phosphorylates Rab8a at T72 (pT72) of its switch 2 α-helix. This post-translational modification enables recruitment of RILPL2, an effector that regulates ciliogenesis in model cell lines. A newly identified RBD motif of RILPL2, termed the X-cap, has been shown to recognize the phosphate via direct interactions between an arginine residue (R132) and pT72 of Rab8a. Here, we show that a second distal arginine (R130) is also essential for phospho-Rab binding by RILPL2. Through structural, biophysical, and cellular studies, we find that R130 stabilizes the primary R132:pT72 salt bridge through favorable enthalpic contributions to the binding affinity. These findings may have implications for the mechanism by which LRRK2 activation leads to assembly of phospho-Rab complexes and subsequent control of their membrane trafficking functions in cells.
Organizational Affiliation:
School of Biochemistry and Immunology, Trinity College Dublin, Dublin, Ireland.