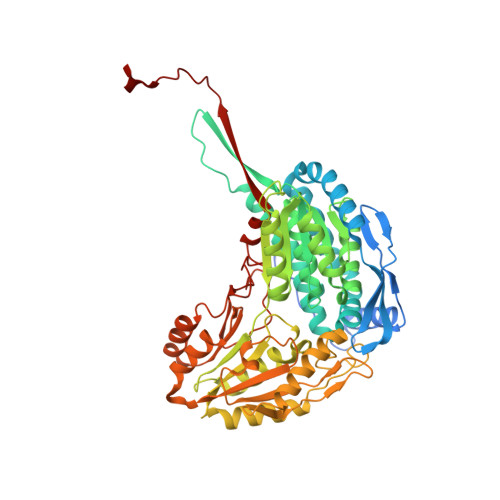
Structural basis for the stereospecific inhibition of the dual proline/hydroxyproline catabolic enzyme ALDH4A1 by trans-4-hydroxy-L-proline.
Bogner, A.N., Stiers, K.M., McKay, C.M., Becker, D.F., Tanner, J.J.(2021) Protein Sci 30: 1714-1722
- PubMed: 34048122
- DOI: https://doi.org/10.1002/pro.4131
- Primary Citation of Related Structures:
7MER, 7MES - PubMed Abstract:
Aldehyde dehydrogenase 4A1 (ALDH4A1) catalyzes the final steps of both proline and hydroxyproline catabolism. It is a dual substrate enzyme that catalyzes the NAD + -dependent oxidations of L-glutamate-γ-semialdehyde to L-glutamate (proline metabolism), and 4-hydroxy-L-glutamate-γ-semialdehyde to 4-erythro-hydroxy-L-glutamate (hydroxyproline metabolism). Here we investigated the inhibition of mouse ALDH4A1 by the six stereoisomers of proline and 4-hydroxyproline using steady-state kinetics and X-ray crystallography. Trans-4-hydroxy-L-proline is the strongest of the inhibitors studied, characterized by a competitive inhibition constant of 0.7 mM, followed by L-proline (1.9 mM). The other compounds are very weak inhibitors (approximately 10 mM or greater). Insight into the selectivity for L-stereoisomers was obtained by solving crystal structures of ALDH4A1 complexed with trans-4-hydroxy-L-proline and trans-4-hydroxy-D-proline. The structures suggest that the 10-fold greater preference for the L-stereoisomer is due to a serine residue that hydrogen bonds to the amine group of trans-4-hydroxy-L-proline. In contrast, the amine group of the D-stereoisomer lacks a direct interaction with the enzyme due to a different orientation of the pyrrolidine ring. These results suggest that hydroxyproline catabolism is subject to substrate inhibition by trans-4-hydroxy-L-proline, analogous to the known inhibition of proline catabolism by L-proline. Also, drugs targeting the first enzyme of hydroxyproline catabolism, by elevating the level of trans-4-hydroxy-L-proline, may inadvertently impair proline catabolism by the inhibition of ALDH4A1.
Organizational Affiliation:
Department of Biochemistry, University of Missouri, Columbia, Missouri, USA.