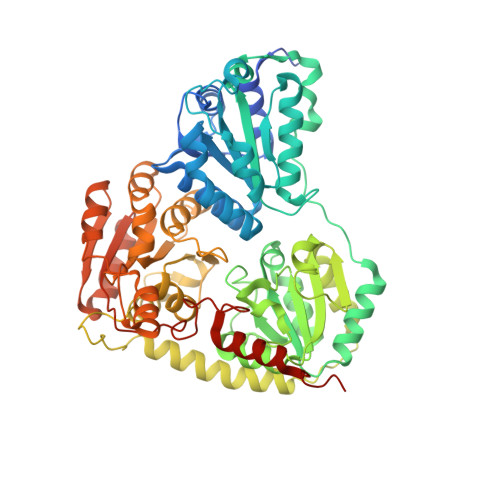
Mechanistic details of the actinobacterial lyase-catalyzed degradation reaction of 2-hydroxyisobutyryl-CoA.
Zahn, M., Konig, G., Pham, H.V.C., Seroka, B., Lazny, R., Yang, G., Ouerfelli, O., Lotowski, Z., Rohwerder, T.(2022) J Biol Chem 298: 101522-101522
- PubMed: 34952003
- DOI: https://doi.org/10.1016/j.jbc.2021.101522
- Primary Citation of Related Structures:
7PT1, 7PT2, 7PT3, 7PT4 - PubMed Abstract:
Actinobacterial 2-hydroxyacyl-CoA lyase reversibly catalyzes the thiamine diphosphate-dependent cleavage of 2-hydroxyisobutyryl-CoA to formyl-CoA and acetone. This enzyme has great potential for use in synthetic one-carbon assimilation pathways for sustainable production of chemicals, but lacks details of substrate binding and reaction mechanism for biochemical reengineering. We determined crystal structures of the tetrameric enzyme in the closed conformation with bound substrate, covalent postcleavage intermediate, and products, shedding light on active site architecture and substrate interactions. Together with molecular dynamics simulations of the covalent precleavage complex, the complete catalytic cycle is structurally portrayed, revealing a proton transfer from the substrate acyl Cβ hydroxyl to residue E493 that returns it subsequently to the postcleavage Cα-carbanion intermediate. Kinetic parameters obtained for mutants E493A, E493Q, and E493K confirm the catalytic role of E493 in the WT enzyme. However, the 10- and 50-fold reduction in lyase activity in the E493A and E493Q mutants, respectively, compared with WT suggests that water molecules may contribute to proton transfer. The putative catalytic glutamate is located on a short α-helix close to the active site. This structural feature appears to be conserved in related lyases, such as human 2-hydroxyacyl-CoA lyase 2. Interestingly, a unique feature of the actinobacterial 2-hydroxyacyl-CoA lyase is a large C-terminal lid domain that, together with active site residues L127 and I492, restricts substrate size to ≤C5 2-hydroxyacyl residues. These details about the catalytic mechanism and determinants of substrate specificity pave the ground for designing tailored catalysts for acyloin condensations for one-carbon and short-chain substrates in biotechnological applications.
Organizational Affiliation:
Centre for Enzyme Innovation, School of Biological Sciences, Institute of Biological and Biomedical Sciences, University of Portsmouth, Portsmouth, United Kingdom. Electronic address: [email protected].