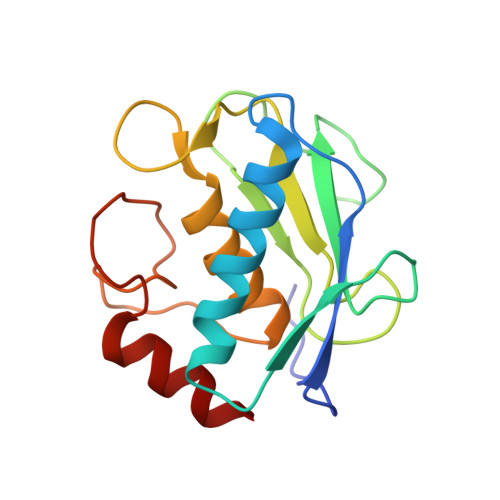
Crystal Structures of MMP-9 Complexes with Five Inhibitors: Contribution of the Flexible Arg424 Side-chain to Selectivity.
Tochowicz, A., Maskos, K., Huber, R., Oltenfreiter, R., Dive, V., Yiotakis, A., Zanda, M., Bode, W., Goettig, P.(2007) J Mol Biol 371: 989-1006
- PubMed: 17599356
- DOI: https://doi.org/10.1016/j.jmb.2007.05.068
- Primary Citation of Related Structures:
2OVX, 2OVZ, 2OW0, 2OW1, 2OW2 - PubMed Abstract:
Human matrix metalloproteinase 9 (MMP-9), also called gelatinase B, is particularly involved in inflammatory processes, bone remodelling and wound healing, but is also implicated in pathological processes such as rheumatoid arthritis, atherosclerosis, tumour growth, and metastasis. We have prepared the inactive E402Q mutant of the truncated catalytic domain of human MMP-9 and co-crystallized it with active site-directed synthetic inhibitors of different binding types. Here, we present the X-ray structures of five MMP-9 complexes with gelatinase-specific, tight binding inhibitors: a phosphinic acid (AM-409), a pyrimidine-2,4,6-trione (RO-206-0222), two carboxylate (An-1 and MJ-24), and a trifluoromethyl hydroxamic acid inhibitor (MS-560). These compounds bind by making a compromise between optimal coordination of the catalytic zinc, favourable hydrogen bond formation in the active-site cleft, and accommodation of their large hydrophobic P1' groups in the slightly flexible S1' cavity, which exhibits distinct rotational conformations of the Pro421 carbonyl group in each complex. In all these structures, the side-chain of Arg424 located at the bottom of the S1' cavity is not defined in the electron density beyond C(gamma), indicating its mobility. However, we suggest that the mobile Arg424 side-chain partially blocks the S1' cavity, which might explain the weaker binding of most inhibitors with a long P1' side-chain for MMP-9 compared with the closely related MMP-2 (gelatinase A), which exhibits a short threonine side-chain at the equivalent position. These novel structural details should facilitate the design of more selective MMP-9 inhibitors.
Organizational Affiliation:
Arbeitsgruppe Proteinaseforschung, Max-Planck-Institut für Biochemie, Am Klopferspitz 18, D-82152 Martinsried, Germany.